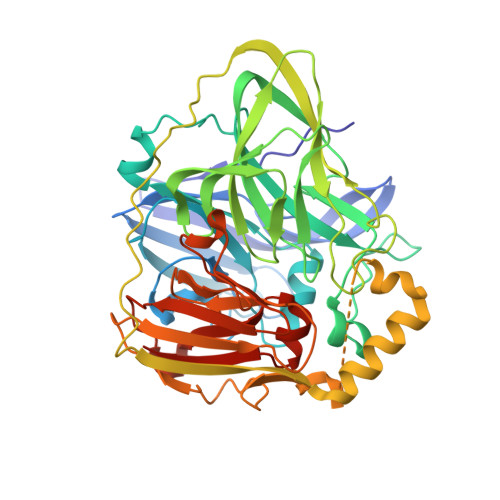
A Labile Regulatory Copper Ion Lies Near the T1 Copper Site in the Multicopper Oxidase CueO.
Roberts, S.A., Wildner, G.F., Grass, G., Weichsel, A., Ambrus, A., Rensing, C., Montfort, W.R.(2003) J Biol Chem 278: 31958-31963
- PubMed: 12794077
- DOI: https://doi.org/10.1074/jbc.M302963200
- Primary Citation of Related Structures:
1N68, 1PF3 - PubMed Abstract:
CueO, a multicopper oxidase, is part of the copper-regulatory cue operon in Escherichia coli, is expressed under conditions of copper stress and shows enhanced oxidase activity when additional copper is present. The 1.7-A resolution structure of a crystal soaked in CuCl2 reveals a Cu(II) ion bound to the protein 7.5 A from the T1 copper site in a region rich in methionine residues. The trigonal bipyramidal coordination sphere is unusual, containing two methionine sulfur atoms, two aspartate carboxylate oxygen atoms, and a water molecule. Asp-439 both ligates the labile copper and hydrogen-bonds to His-443, which ligates the T1 copper. This arrangement may mediate electron transfer from substrates to the T1 copper. Mutation of residues bound to the labile copper results in loss of oxidase activity and of copper tolerance, confirming a regulatory role for this site. The methionine-rich portion of the protein, which is similar to that of other proteins involved in copper homeostasis, does not display additional copper binding. The type 3 copper atoms of the trinuclear cluster in the structure are bridged by a chloride ion that completes a square planar coordination sphere for the T2 copper atom but does not affect oxidase activity.
Organizational Affiliation:
Departments of Biochemistry and Molecular Biophysics and Soil, Water, and Environmental Science, University of Arizona, Tucson, Arizona 85721.