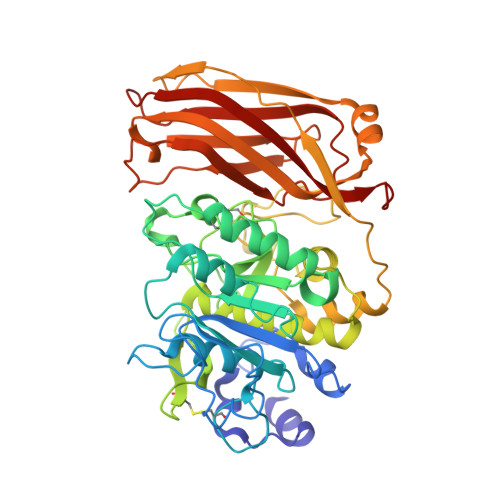
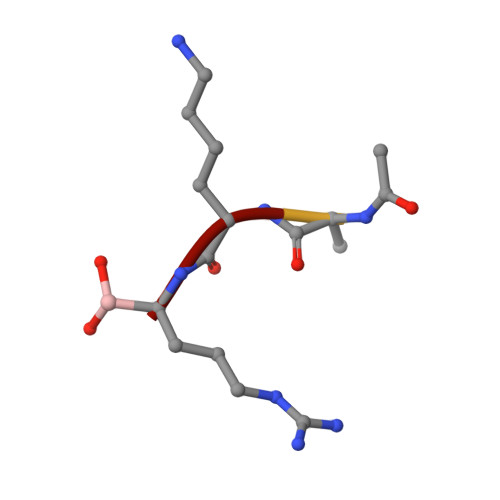
2.4 A Resolution Crystal Structure of the Prototypical Hormone-Processing Protease Kex2 in Complex with an Ala-Lys-Arg Boronic Acid Inhibitor
Holyoak, T., Wilson, M.A., Fenn, T.D., Kettner, C.A., Petsko, G.A., Fuller, R.S., Ringe, D.(2003) Biochemistry 42: 6709-6718
- PubMed: 12779325
- DOI: https://doi.org/10.1021/bi034434t
- Primary Citation of Related Structures:
1OT5 - PubMed Abstract:
This paper reports the first structure of a member of the Kex2/furin family of eukaryotic pro-protein processing proteases, which cleave sites consisting of pairs or clusters of basic residues. Reported is the 2.4 A resolution crystal structure of the two-domain protein ssKex2 in complex with an Ac-Ala-Lys-boroArg inhibitor (R = 20.9%, R(free) = 24.5%). The Kex2 proteolytic domain is similar in its global fold to the subtilisin-like superfamily of degradative proteases. Analysis of the complex provides a structural basis for the extreme selectivity of this enzyme family that has evolved from a nonspecific subtilisin-like ancestor. The P-domain of ssKex2 has a novel jelly roll like fold consisting of nine beta strands and may potentially be involved, along with the buried Ca(2+) ion, in creating the highly determined binding site for P(1) arginine.
Organizational Affiliation:
Rosenstiel Basic Medical Sciences Research Center, Brandeis University, Waltham, Massachusetts 02454, USA.