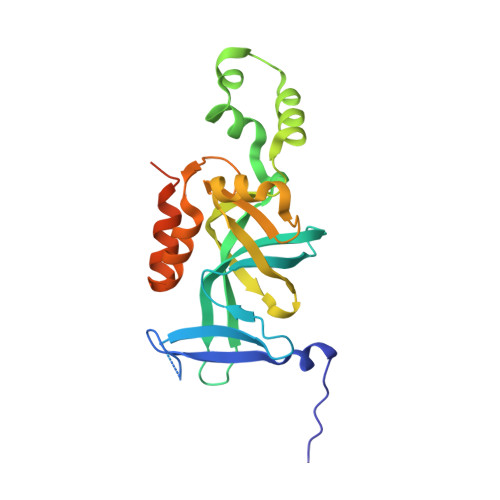
Structure and function of the BAH-containing domain of Orc1p in epigenetic silencing.
Zhang, Z., Hayashi, M.K., Merkel, O., Stillman, B., Xu, R.M.(2002) EMBO J 21: 4600-4611
- PubMed: 12198162
- DOI: https://doi.org/10.1093/emboj/cdf468
- Primary Citation of Related Structures:
1M4Z - PubMed Abstract:
The N-terminal domain of the largest subunit of the Saccharomyces cerevisiae origin recognition complex (Orc1p) functions in transcriptional silencing and contains a bromo-adjacent homology (BAH) domain found in some chromatin-associated proteins including Sir3p. The 2.2 A crystal structure of the N-terminal domain of Orc1p revealed a BAH core and a non-conserved helical sub-domain. Mutational analyses demonstrated that the helical sub-domain was necessary and sufficient to bind Sir1p, and critical for targeting Sir1p primarily to the cis-acting E silencers at the HMR and HML silent chromatin domains. In the absence of the BAH domain, approximately 14-20% of cells in a population were silenced at the HML locus. Moreover, the distributions of the Sir2p, Sir3p and Sir4p proteins, while normal, were at levels lower than found in wild-type cells. Thus, in the absence of the Orc1p BAH domain, HML resembled silencing of genes adjacent to telomeres. These data are consistent with the view that the Orc1p-Sir1p interaction at the E silencers ensures stable inheritance of pre-established Sir2p, Sir3p and Sir4p complexes at the silent mating type loci.
Organizational Affiliation:
Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724, USA.