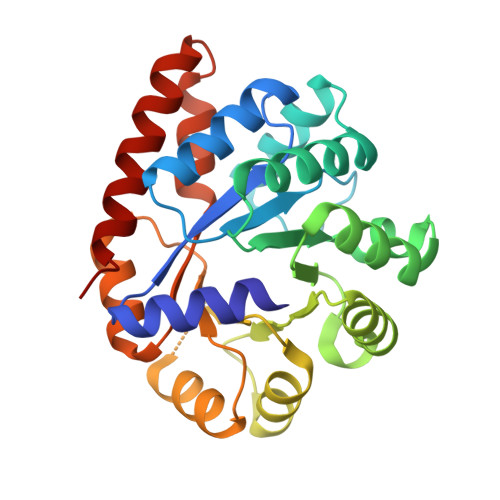
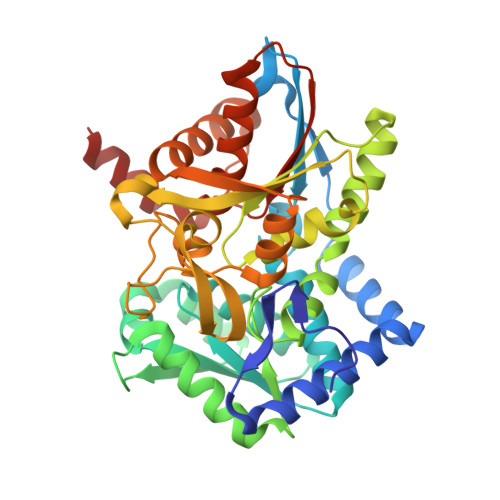
On the Role of alphaTHR183 in the Allosteric Regulation and Catalytic Mechanism of Tryptophan Synthase
Kulik, V., Weyand, M., Siedel, R., Niks, D., Arac, D., Dunn, M.F., Schlichting, I.(2002) J Mol Biol 324: 677-690
- PubMed: 12460570
- DOI: https://doi.org/10.1016/s0022-2836(02)01109-9
- Primary Citation of Related Structures:
1K8X, 1KFB, 1KFC, 1KFE, 1KFJ, 1KFK - PubMed Abstract:
The catalytic activity and substrate channeling of the pyridoxal 5'-phosphate-dependent tryptophan synthase alpha(2)beta(2) complex is regulated by allosteric interactions that modulate the switching of the enzyme between open, low activity and closed, high activity states during the catalytic cycle. The highly conserved alphaThr183 residue is part of loop alphaL6 and is located next to the alpha-active site and forms part of the alpha-beta subunit interface. The role of the interactions of alphaThr183 in alpha-site catalysis and allosteric regulation was investigated by analyzing the kinetics and crystal structures of the isosteric mutant alphaThr183Val. The mutant displays strongly impaired allosteric alpha-beta communication, and the catalytic activity of the alpha-reaction is reduced one hundred fold, whereas the beta-activity is not affected. The structural work establishes that the basis for the missing inter-subunit signaling is the lack of loop alphaL6 closure even in the presence of the alpha-subunit ligands, 3-indolyl-D-glycerol 3'-phosphate, or 3-indolylpropanol 3'-phosphate. The structural basis for the reduced alpha-activity has its origins in the missing hydrogen bond between alphaThr183 and the catalytic residue, alphaAsp60.
Organizational Affiliation:
Max Planck Institut für Molekulare Physiologie, Abeilung für Biophysikalische Chemie, Otto Hahn Str 11, 44227 Dortmund, Germany.