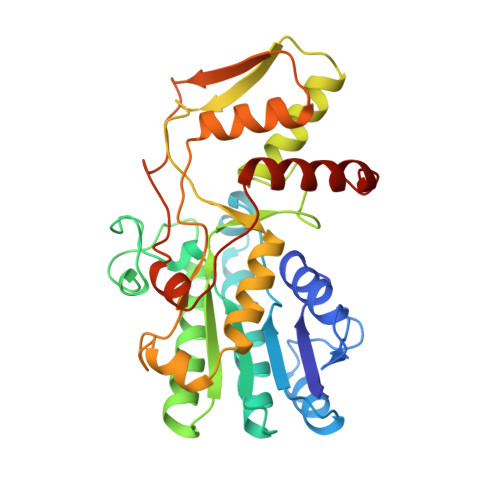
GDP-fucose synthetase from Escherichia coli: structure of a unique member of the short-chain dehydrogenase/reductase family that catalyzes two distinct reactions at the same active site.
Somers, W.S., Stahl, M.L., Sullivan, F.X.(1998) Structure 6: 1601-1612
- PubMed: 9862812
- DOI: https://doi.org/10.1016/s0969-2126(98)00157-9
- Primary Citation of Related Structures:
1BSV, 1FXS, 1GFS - PubMed Abstract:
. In all species examined, GDP-fucose is synthesized from GDP-mannose in a three-step reaction catalyzed by two enzymes, GDP-mannose 4,6 dehydratase and a dual function 3, 5-epimerase-4-reductase named GDP-fucose synthetase. In this latter aspect fucose biosynthesis differs from that of other deoxy and dideoxy sugars, in which the epimerase and reductase activities are present as separate enzymes. Defects in GDP-fucose biosynthesis have been shown to affect nodulation in bacteria, stem development in plants, and are associated with the immune defect leukocyte adhesion deficiency type II in humans.
Organizational Affiliation:
Small Molecule Drug Discovery Genetics Institute, Inc. 87 Cambridgepark Drive, Cambridge, MA 02140, USA.