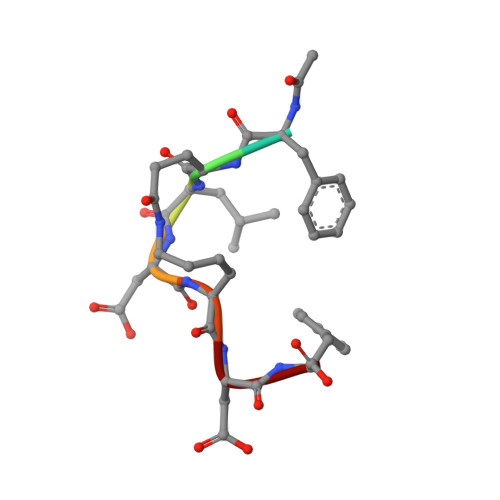
Structure-based optimization of peptide inhibitors of mammalian ribonucleotide reductase.
Pellegrini, M., Liehr, S., Fisher, A.L., Laub, P.B., Cooperman, B.S., Mierke, D.F.(2000) Biochemistry 39: 12210-12215
- PubMed: 11015199
- DOI: https://doi.org/10.1021/bi001323a
- Primary Citation of Related Structures:
1FOZ - PubMed Abstract:
Mammalian ribonucleotide reductase (mRR), a potential target for cancer intervention, is composed of two subunits, mR1 and mR2, whose association is critical for enzyme activity. In this article we describe the structural features of the mRR-inhibitor Ac-F-c[ELAK]-DF (Peptide 3) while bound to the mR1 subunit as determined by transferred NOEs. Peptide 3 is a cyclic analogue of the N-acetylated form of the heptapeptide C-terminus of the mR2 subunit (Ac-FTLDADF), which is the link between the two subunits and previously shown to be the minimal sequence inhibitor mRR by competing with mR2 for binding to mR1. Structural refinement employing an ensemble-based, full-relaxation matrix approach resulted in two structures varying in the conformations of F(1) and the cyclic lactam side chains of E(2) and K(5). The remainder of the molecule, both backbone and side chains, is extremely well-defined, with an RMSD of 0.54 A. The structural features of this conformationally constrained analogue provide unique insight into the requirements for binding to mR1, critical for further inhibitor development.
Organizational Affiliation:
Department of Molecular Pharmacology, Division of Biology and Medicine, and Department of Chemistry, Brown University, Providence, Rhode Island 02912, USA.