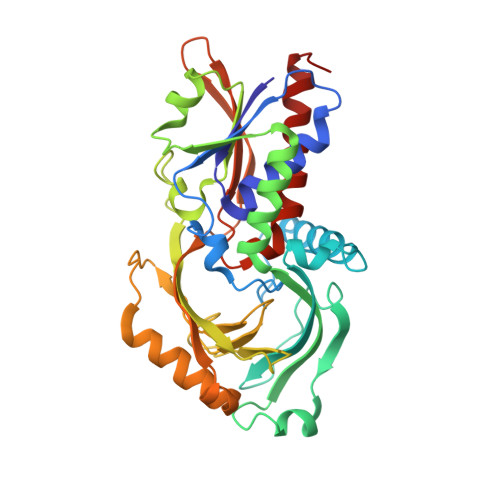
Three-dimensional structure of the purple intermediate of porcine kidney D-amino acid oxidase. Optimization of the oxidative half-reaction through alignment of the product with reduced flavin.
Mizutani, H., Miyahara, I., Hirotsu, K., Nishina, Y., Shiga, K., Setoyama, C., Miura, R.(2000) J Biochem 128: 73-81
- PubMed: 10876160
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a022732
- Primary Citation of Related Structures:
1EVI - PubMed Abstract:
The three-dimensional structure of the purple intermediate of porcine kidney D-amino acid oxidase (DAO) was solved by cryo-X-ray crystallography; the purple intermediate is known to comprise a complex between the dehydrogenated product, an imino acid, and the reduced form of DAO. The crystalline purple intermediate was obtained by anaerobically soaking crystals of oxidized DAO in a buffer containing excess D-proline as the substrate. The dehydrogenated product, delta(1)-pyrrolidine-2-carboxylate (DPC), is found sandwiched between the phenol ring of Tyr 224 and the planar reduced flavin ring. The cationic protonated imino nitrogen is within hydrogen-bonding distance of the backbone carbonyl oxygen of Gly 313. The carboxyl group of DPC is recognized by the Arg 283 guanidino and Tyr 228 hydroxyl groups through ion-pairing and hydrogen-bonding, respectively. The (+)HN=C double bond of DPC overlaps the N(5)-C(4a) bond of reduced flavin. The electrostatic effect of the cationic nitrogen of DPC is suggested to shift the resonance hybridization of anionic reduced flavin toward a canonical form with a negative charge at C(4a), thereby augmenting the electron density at C(4a), from which electrons are transferred to molecular oxygen during reoxidation of reduced flavin. The reactivity of reduced flavin in the purple intermediate, therefore, is enhanced through the alignment of DPC with respect to reduced flavin.
Organizational Affiliation:
Department of Chemistry, Faculty of Science, Osaka City University, Sugimoto, Sumiyoshi-ku, Osaka 558-8585, Japan.