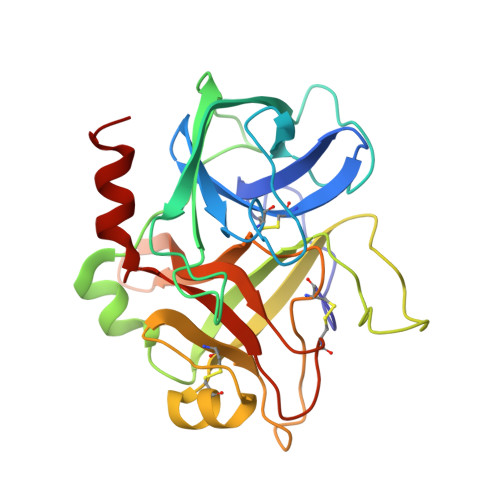
Crystallographic determination of the structures of human alpha-thrombin complexed with BMS-186282 and BMS-189090.
Malley, M.F., Tabernero, L., Chang, C.Y., Ohringer, S.L., Roberts, D.G., Das, J., Sack, J.S.(1996) Protein Sci 5: 221-228
- PubMed: 8745399
- DOI: https://doi.org/10.1002/pro.5560050205
- Primary Citation of Related Structures:
1BMM, 1BMN - PubMed Abstract:
The crystallographic structures of the ternary complexes of human alpha-thrombin with hirugen (a sulfated hirudin fragment) and the small-molecule active site thrombin inhibitors BMS-186282 and BMS-189090 have been determined at 2.6 and 2.8 A. In both cases, the inhibitors, which adopt very similar bound conformations, bind in an antiparallel beta-strand arrangement relative to the thrombin main chain in a manner like that reported for PPACK, D-Phe-Pro-Arg-CH2Cl. They do, however, exhibit differences in the binding of the alkyl guanidine moiety in the specificity pocket. Numerous hydrophilic and hydrophobic interactions serve to stabilize the inhibitors in the binding pocket. Although PPACK forms covalent bonds to both serine and the histidine of the catalytic triad of thrombin, neither BMS-186282 nor BMS-189090 bind covalently and only BMS-186282 forms a hydrogen bond to the serine of the catalytic triad. Both inhibitors bind with high affinity (Ki = 79 nM and 3.6 nM, respectively) and are highly selective for thrombin over trypsin and other serine proteases.
Organizational Affiliation:
Department of Solid State Chemistry, Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, New Jersey 08543-4000, USA. malley@bms.com