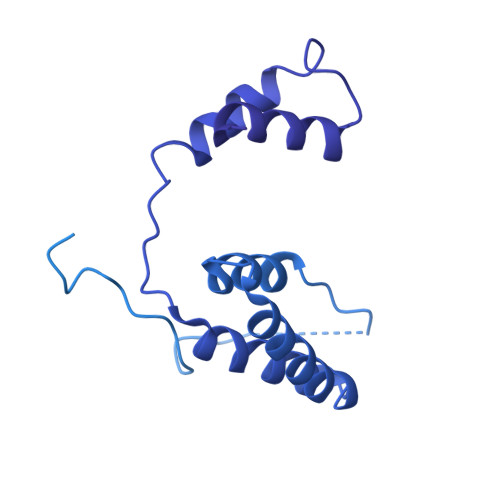
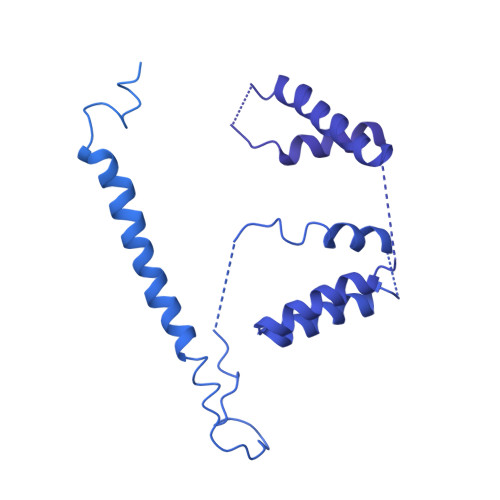
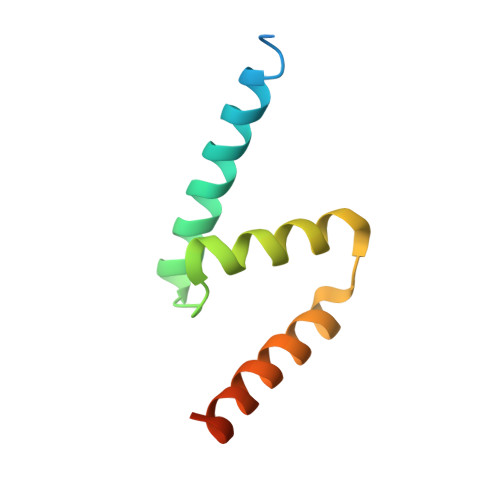
MZT Proteins Form Multi-Faceted Structural Modules in the gamma-Tubulin Ring Complex.
Wieczorek, M., Huang, T.L., Urnavicius, L., Hsia, K.C., Kapoor, T.M.(2020) Cell Rep 31: 107791-107791
- PubMed: 32610146
- DOI: https://doi.org/10.1016/j.celrep.2020.107791
- Primary Citation of Related Structures:
6M33, 6X0U, 6X0V - PubMed Abstract:
Microtubule organization depends on the γ-tubulin ring complex (γ-TuRC), a ∼2.3-MDa nucleation factor comprising an asymmetric assembly of γ-tubulin and GCP2-GCP6. However, it is currently unclear how the γ-TuRC-associated microproteins MZT1 and MZT2 contribute to the structure and regulation of the holocomplex. Here, we report cryo-EM structures of MZT1 and MZT2 in the context of the native human γ-TuRC. MZT1 forms two subcomplexes with the N-terminal α-helical domains of GCP3 or GCP6 (GCP-NHDs) within the γ-TuRC "lumenal bridge." We determine the X-ray structure of recombinant MZT1/GCP6-NHD and find it is similar to that within the native γ-TuRC. We identify two additional MZT/GCP-NHD-like subcomplexes, one of which is located on the outer face of the γ-TuRC and comprises MZT2 and GCP2-NHD in complex with a centrosomin motif 1 (CM1)-containing peptide. Our data reveal how MZT1 and MZT2 establish multi-faceted, structurally mimetic "modules" that can expand structural and regulatory interfaces in the γ-TuRC.
Organizational Affiliation:
Laboratory of Chemistry and Cell Biology, The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.