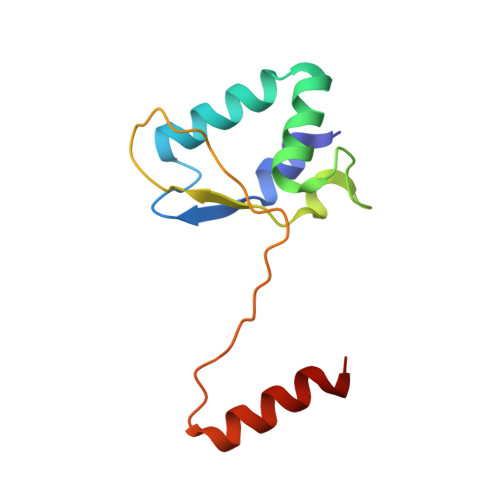
Crystal structure of the catalytic C-lobe of the HECT-type ubiquitin ligase E6AP.
Ries, L.K., Liess, A.K.L., Feiler, C.G., Spratt, D.E., Lowe, E.D., Lorenz, S.(2020) Protein Sci 29: 1550-1554
- PubMed: 31994269
- DOI: https://doi.org/10.1002/pro.3832
- Primary Citation of Related Structures:
6TGK - PubMed Abstract:
The HECT-type ubiquitin ligase E6AP (UBE3A) is critically involved in several neurodevelopmental disorders and human papilloma virus-induced cervical tumorigenesis; the structural mechanisms underlying the activity of this crucial ligase, however, are incompletely understood. Here, we report a crystal structure of the C-terminal lobe ("C-lobe") of the catalytic domain of E6AP that reveals two molecules in a domain-swapped, dimeric arrangement. Interestingly, the molecular hinge that enables this structural reorganization with respect to the monomeric fold coincides with the active-site region. While such dimerization is unlikely to occur in the context of full-length E6AP, we noticed a similar domain swap in a crystal structure of the isolated C-lobe of another HECT-type ubiquitin ligase, HERC6. This may point to conformational strain in the active-site region of HECT-type ligases with possible implications for catalysis. SIGNIFICANCE STATEMENT: The HECT-type ubiquitin ligase E6AP has key roles in human papilloma virus-induced cervical tumorigenesis and certain neurodevelopmental disorders. Here, we present a crystal structure of the C-terminal, catalytic lobe of E6AP, providing basic insight into the conformational properties of this functionally critical region of HECT-type ligases.
Organizational Affiliation:
Rudolf Virchow Center for Experimental Biomedicine, University of Würzburg, Würzburg, Germany.