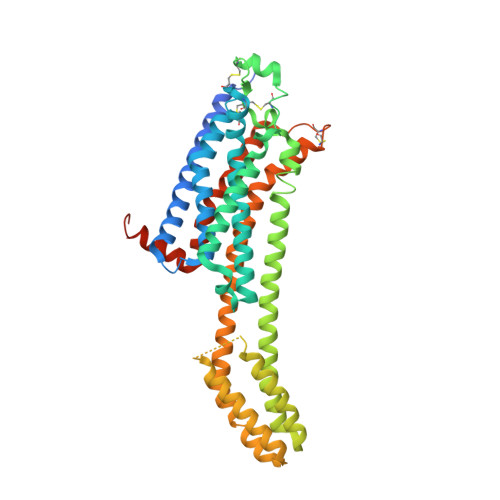
High-viscosity injector-based pink-beam serial crystallography of microcrystals at a synchrotron radiation source.
Martin-Garcia, J.M., Zhu, L., Mendez, D., Lee, M.Y., Chun, E., Li, C., Hu, H., Subramanian, G., Kissick, D., Ogata, C., Henning, R., Ishchenko, A., Dobson, Z., Zhang, S., Weierstall, U., Spence, J.C.H., Fromme, P., Zatsepin, N.A., Fischetti, R.F., Cherezov, V., Liu, W.(2019) IUCrJ 6: 412-425
- PubMed: 31098022
- DOI: https://doi.org/10.1107/S205225251900263X
- Primary Citation of Related Structures:
6MH6, 6MH8 - PubMed Abstract:
Since the first successful serial crystallography (SX) experiment at a synchrotron radiation source, the popularity of this approach has continued to grow showing that third-generation synchrotrons can be viable alternatives to scarce X-ray free-electron laser sources. Synchrotron radiation flux may be increased ∼100 times by a moderate increase in the bandwidth ('pink beam' conditions) at some cost to data analysis complexity. Here, we report the first high-viscosity injector-based pink-beam SX experiments. The structures of proteinase K (PK) and A 2A adenosine receptor (A 2A AR) were determined to resolutions of 1.8 and 4.2 Å using 4 and 24 consecutive 100 ps X-ray pulse exposures, respectively. Strong PK data were processed using existing Laue approaches, while weaker A 2A AR data required an alternative data-processing strategy. This demonstration of the feasibility presents new opportunities for time-resolved experiments with microcrystals to study structural changes in real time at pink-beam synchrotron beamlines worldwide.
Organizational Affiliation:
Biodesign Center for Applied Structural Discovery, Biodesign Institute, Arizona State University, 727 East Tyler Street, Tempe, AZ 85287, USA.