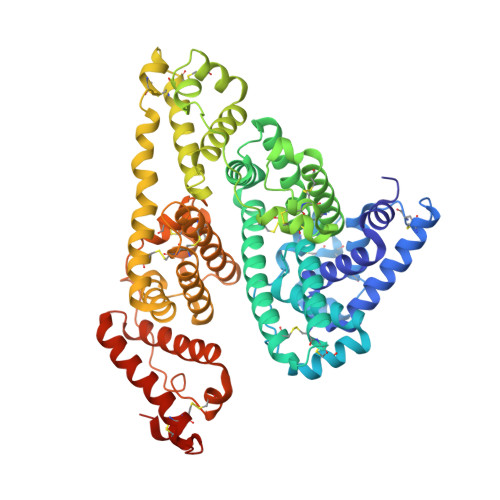
Crystal structure analysis of human serum albumin complexed with sodium 4-phenylbutyrate.
Kawai, A., Yamasaki, K., Enokida, T., Miyamoto, S., Otagiri, M.(2018) Biochem Biophys Rep 13: 78-82
- PubMed: 29387812
- DOI: https://doi.org/10.1016/j.bbrep.2018.01.006
- Primary Citation of Related Structures:
5YOQ - PubMed Abstract:
Sodium 4-phenylbutyrate (PB) is an orphan drug for the treatment of urea cycle disorders. It also inhibits the development of endoplasmic reticulum stress, the action of histone deacetylases and as a regulator of the hepatocanalicular transporter. PB is generally considered to have the potential for use in the treatment of the diseases such as cancer, neurodegenerative diseases and metabolic diseases. In a previous study, we reported that PB is primarily bound to human serum albumin (HSA) in plasma and its binding site is drug site 2. However, details of the binding mode of PB to HSA remain unknown. To address this issue, we examined the crystal structure of HSA with PB bound to it. The structure of the HSA-PB complex indicates that the binding mode of PB to HSA is quite similar to that for octanoate or drugs that bind to drug site 2, as opposed to that for other medium-chain length of fatty acids. These findings provide useful basic information related to drug-HSA interactions. Moreover, the information presented herein is valuable in terms of providing safe and efficient treatment and diagnosis in clinical settings.
Organizational Affiliation:
Faculty of Pharmaceutical Sciences, Sojo University, 4-22-1 Ikeda, Nishi-ku, Kumamoto 860-0082, Japan.