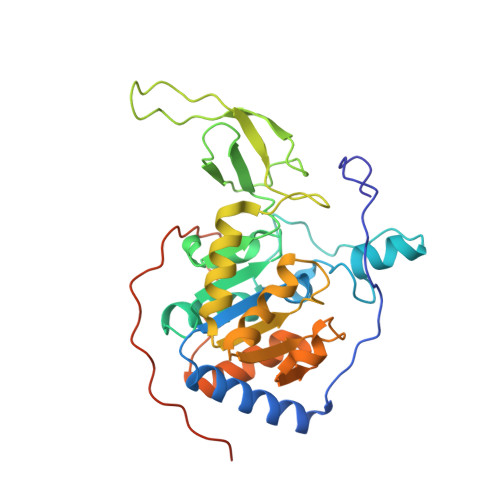
Identification of a cellularly active SIRT6 allosteric activator.
Huang, Z., Zhao, J., Deng, W., Chen, Y., Shang, J., Song, K., Zhang, L., Wang, C., Lu, S., Yang, X., He, B., Min, J., Hu, H., Tan, M., Xu, J., Zhang, Q., Zhong, J., Sun, X., Mao, Z., Lin, H., Xiao, M., Chin, Y.E., Jiang, H., Xu, Y., Chen, G., Zhang, J.(2018) Nat Chem Biol 14: 1118-1126
- PubMed: 30374165
- DOI: https://doi.org/10.1038/s41589-018-0150-0
- Primary Citation of Related Structures:
5Y2F - PubMed Abstract:
SIRT6, a member of the SIRT deacetylase family, is responsible for deacetylation of histone H3 N ε -acetyl-lysines 9 (H3K9ac) and 56 (H3K56ac). As a tumor suppressor, SIRT6 has frequently been found to have low expression in various cancers. Here, we report the identification of MDL-800, a selective SIRT6 activator. MDL-800 increased the deacetylase activity of SIRT6 by up to 22-fold via binding to an allosteric site; this interaction led to a global decrease in H3K9ac and H3K56ac levels in human hepatocellular carcinoma (HCC) cells. Consequently, MDL-800 inhibited the proliferation of HCC cells via SIRT6-driven cell-cycle arrest and was effective in a tumor xenograft model. Together, these data demonstrate that pharmacological activation of SIRT6 is a potential therapeutic approach for the treatment of HCC. MDL-800 is a first-in-class small-molecule cellular SIRT6 activator that can be used to physiologically and pathologically investigate the roles of SIRT6 deacetylation.
Organizational Affiliation:
Key Laboratory of Cell Differentiation and Apoptosis, Ministry of Education, Department of Pathophysiology, Ruijin Hospital, Shanghai Jiao-Tong University School of Medicine, Shanghai, China.