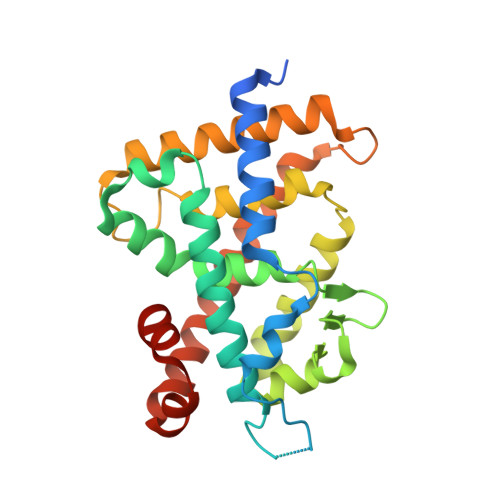
Vitamin D Analogues with a p-Hydroxyphenyl Group at the C25 Position: Crystal Structure of Vitamin D Receptor Ligand-Binding Domain Complexed with the Ligand Explains the Mechanism Underlying Full Antagonistic Action
Kato, A., Yamao, M., Hashihara, Y., Ishida, H., Itoh, T., Yamamoto, K.(2017) J Med Chem 60: 8394-8406
- PubMed: 28954197
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00819
- Primary Citation of Related Structures:
5XPL, 5XPM, 5XPN, 5XPO, 5XPP - PubMed Abstract:
Vitamin D receptor (VDR) antagonists can be classified into two categories: the first category of VDR antagonists, which do not stabilize the helix 11-12, and the second category of antagonists, which destabilize the helix 6-7 region. To elucidate the mechanism underlying the first category antagonists by using the crystal structure, we designed and synthesized several VDR ligands with a p-hydroxyphenyl group at the C25-position. Of these, 22S-butyl-25-carbonyl analogue 5b and 25-di-p-hydoroxyphenyl analogues 6a,b showed strong antagonistic activity. We succeeded in cocrystallizing the ligand-binding domain of VDR complexed with 5b and found that the structure showed an alternative conformation of the helix 11-12 that explained the mechanism of the first category antagonists. Taking the present and previous studies together, we could elucidate the mechanisms underlying first and second categories antagonists based on individual crystal structures. This study provides significant insights into antagonism against not only VDR but also nuclear receptors.
Organizational Affiliation:
Laboratory of Drug Design and Medicinal Chemistry, Showa Pharmaceutical University , 3-3165 Higashi-Tamagawagakuen, Machida, Tokyo 194-8543, Japan.