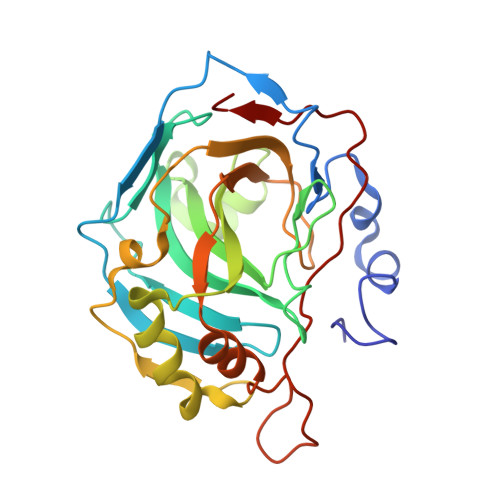
Structural study of the location of the phenyl tail of benzene sulfonamides and the effect on human carbonic anhydrase inhibition.
Guzel-Akdemir, O., Biswas, S., Lastra, K., McKenna, R., Supuran, C.T.(2013) Bioorg Med Chem 21: 6674-6680
- PubMed: 24012377
- DOI: https://doi.org/10.1016/j.bmc.2013.08.011
- Primary Citation of Related Structures:
4ITP - PubMed Abstract:
The crystal structure of 4-phenylacetamidomethyl-benzenesulfonamide (4ITP) bound to human carbonic anhydrase (hCA, EC 4.2.1.1) II is reported. 4ITP is a medium potency hCA I and II inhibitor (KIs of 54-75nM), a strong mitochondrial CA VA/VB inhibitor (KIs of 8.3-8.6nM) and a weak transmembrane CA inhibitor (KIs of 136-212nM against hCA IX and XII). This elongated compound binds in an extended conformation to hCA II, with its tail lying towards the hydrophobic half of the active site whereas the sulfonamide moiety coordinates the zinc ion. The present structure was compared to that of structurally related aromatic sulfonamides, such as 4-phenylacetamido-benzene-sulfonamide (3OYS), 4-(2-mercaptophenylacetamido)-benzene-sulfonamide (2HD6) and 4-(3-nitrophenyl)-ureido-benzenesulfonamide (3N2P). Homology models of the hCA I, VA, VB, IX and XII structures were build which afforded an understanding of the amino acids involved in the binding of these compounds to these isoforms. The main conclusion of the study is that the orientation of the tail moiety and the presence of flexible linkers as well polar groups in it, strongly influence the potency and the selectivity of the sulfonamides for the inhibition of cytosolic, mitochondrial or transmembrane CA isoforms.
Organizational Affiliation:
Università degli Studi di Firenze, Polo Scientifico, Laboratorio di Chimica Bioinorganica, Rm. 188, Via della Lastruccia 3, 50019 Sesto Fiorentino, Italy; Istanbul University, Faculty of Pharmacy, Department of Pharmaceutical Chemistry, 34116 Beyazit, Istanbul, Turkey.