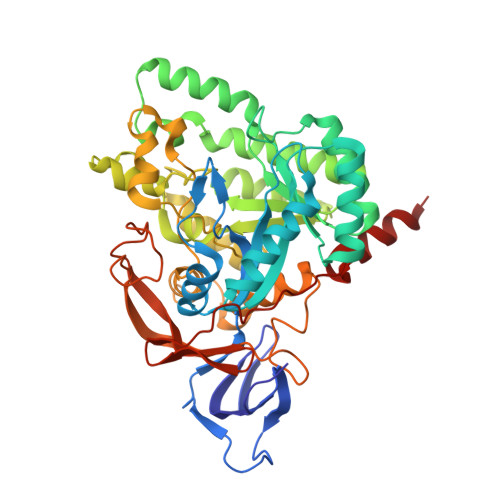
Crystal Structure of Human Crmp-4: Correction of Intensities for Lattice-Translocation Disorder
Ponnusamy, R., Lebedev, A., Pahlow, S., Lohkamp, B.(2014) Acta Crystallogr D Biol Crystallogr 70: 1680
- PubMed: 24914979
- DOI: https://doi.org/10.1107/S1399004714006634
- Primary Citation of Related Structures:
4CNS, 4CNT, 4CNU - PubMed Abstract:
Collapsin response mediator proteins (CRMPs) are cytosolic phosphoproteins that are mainly involved in neuronal cell development. In humans, the CRMP family comprises five members. Here, crystal structures of human CRMP-4 in a truncated and a full-length version are presented. The latter was determined from two types of crystals, which were either twinned or partially disordered. The crystal disorder was coupled with translational NCS in ordered domains and manifested itself with a rather sophisticated modulation of intensities. The data were demodulated using either the two-lattice treatment of lattice-translocation effects or a novel method in which demodulation was achieved by independent scaling of several groups of intensities. This iterative protocol does not rely on any particular parameterization of the modulation coefficients, but uses the current refined structure as a reference. The best results in terms of R factors and map correlation coefficients were obtained using this new method. The determined structures of CRMP-4 are similar to those of other CRMPs. Structural comparison allowed the confirmation of known residues, as well as the identification of new residues, that are important for the homo- and hetero-oligomerization of these proteins, which are critical to nerve-cell development. The structures provide further insight into the effects of medically relevant mutations of the DPYSL-3 gene encoding CRMP-4 and the putative enzymatic activities of CRMPs.
Organizational Affiliation:
Instituto de Technologia Química e Biológica, Universidade Nova de Lisboa, Avenida da República, EAN, 2781-901 Oeiras, Portugal.