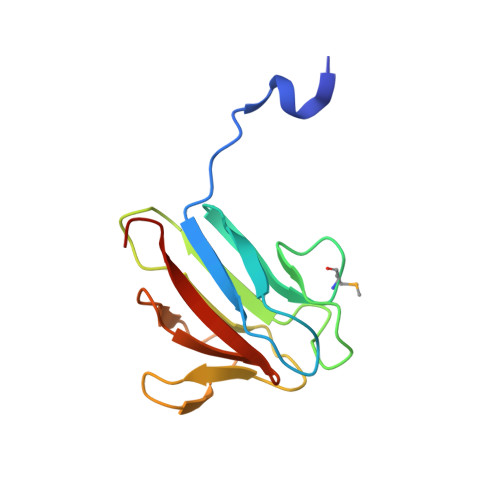
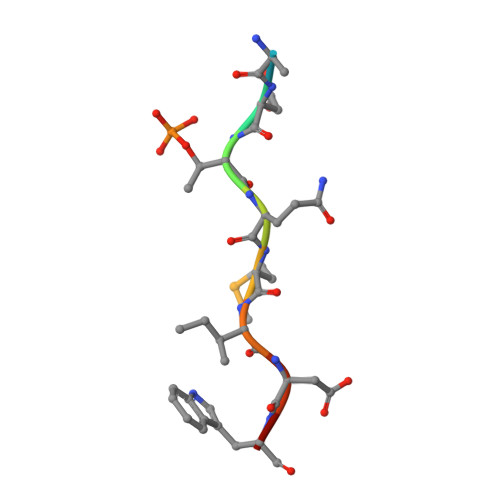
The molecular basis of ATM-dependent dimerization of the Mdc1 DNA damage checkpoint mediator.
Jungmichel, S., Clapperton, J.A., Lloyd, J., Hari, F.J., Spycher, C., Pavic, L., Li, J., Haire, L.F., Bonalli, M., Larsen, D.H., Lukas, C., Lukas, J., MacMillan, D., Nielsen, M.L., Stucki, M., Smerdon, S.J.(2012) Nucleic Acids Res 40: 3913-3928
- PubMed: 22234878
- DOI: https://doi.org/10.1093/nar/gkr1300
- Primary Citation of Related Structures:
3UN0, 3UOT - PubMed Abstract:
Mdc1 is a large modular phosphoprotein scaffold that maintains signaling and repair complexes at double-stranded DNA break sites. Mdc1 is anchored to damaged chromatin through interaction of its C-terminal BRCT-repeat domain with the tail of γH2AX following DNA damage, but the role of the N-terminal forkhead-associated (FHA) domain remains unclear. We show that a major binding target of the Mdc1 FHA domain is a previously unidentified DNA damage and ATM-dependent phosphorylation site near the N-terminus of Mdc1 itself. Binding to this motif stabilizes a weak self-association of the FHA domain to form a tight dimer. X-ray structures of free and complexed Mdc1 FHA domain reveal a 'head-to-tail' dimerization mechanism that is closely related to that seen in pre-activated forms of the Chk2 DNA damage kinase, and which both positively and negatively influences Mdc1 FHA domain-mediated interactions in human cells prior to and following DNA damage.
Organizational Affiliation:
Institute of Veterinary Biochemistry and Molecular Biology, University of Zürich - Irchel, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland.