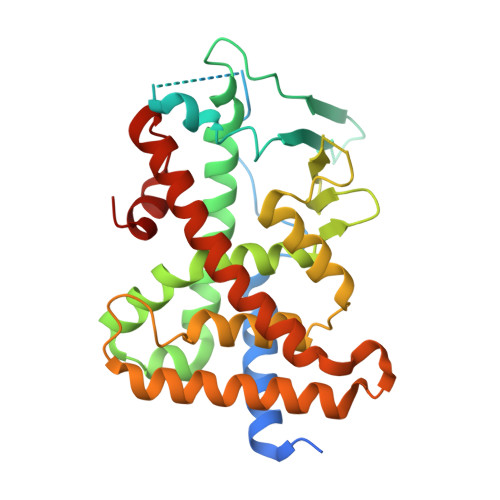
Activation of the human nuclear xenobiotic receptor PXR by the reverse transcriptase-targeted anti-HIV drug PNU-142721.
Cheng, Y., Redinbo, M.R.(2011) Protein Sci 20: 1713-1719
- PubMed: 21805522
- DOI: https://doi.org/10.1002/pro.706
- Primary Citation of Related Structures:
3R8D - PubMed Abstract:
The human pregnane X receptor (PXR) is a member of the nuclear receptor superfamily of ligand-regulated transcription factors. PXR responds to a structurally diverse variety of endogenous and xenobiotic compounds, and coordinates the expression of genes central to the metabolism and excretion of potentially harmful chemicals, including human therapeutics. The reverse transcriptase inhibitor PNU-142721 has been designed to treat human immunodeficiency virus (HIV) infection. Although this compound has anti-HIV activity, it was established using cell-based assays that PNU-142721 is an efficacious PXR agonist. We present here the 2.8 Å resolution crystal structure of the human PXR ligand-binding domain in complex with PNU-142721. PXR employs one hydrogen bond and fourteen van der Waals contacts to interact with the ligand, but allows two loops adjacent to the ligand-binding pocket to remain disordered in the structure. These observations highlight the role structural flexibility plays in PXR's promiscuous responses to xenobiotics. The crystal structure also explains why PNU-173575, a thiomethyl metabolite of PNU-142721, exhibits enhanced PXR activation relative to the unmodified compound and why PNU-142721 can also activate rat PXR. Taken together, the results presented here elucidate the structural basis for PXR activation by PNU-142721 and related chemicals.
Organizational Affiliation:
Department of Biochemistry & Biophysics, University of North Carolina at Chapel Hill, North Carolina 27599-3290, USA.