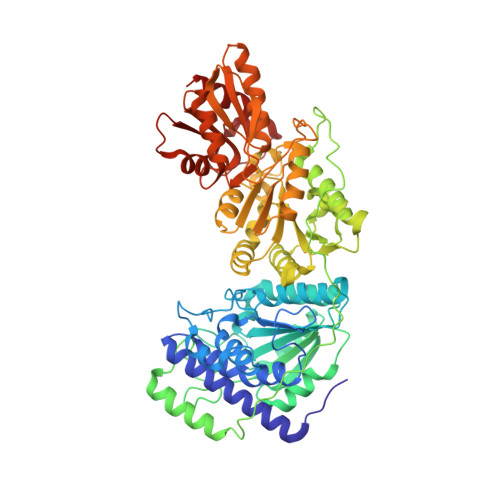
The crystal structure of human transketolase and new insights into its mode of action.
Mitschke, L., Parthier, C., Schroder-Tittmann, K., Coy, J., Ludtke, S., Tittmann, K.(2010) J Biol Chem 285: 31559-31570
- PubMed: 20667822
- DOI: https://doi.org/10.1074/jbc.M110.149955
- Primary Citation of Related Structures:
3MOS - PubMed Abstract:
The crystal structure of human transketolase (TKT), a thiamine diphosphate (ThDP) and Ca(2+)-dependent enzyme that catalyzes the interketol transfer between ketoses and aldoses as part of the pentose phosphate pathway, has been determined to 1.75 Å resolution. The recombinantly produced protein crystallized in space group C2 containing one monomer in the asymmetric unit. Two monomers form the homodimeric biological assembly with two identical active sites at the dimer interface. Although the protomer exhibits the typical three (α/β)-domain structure and topology reported for TKTs from other species, structural differences are observed for several loop regions and the linker that connects the PP and Pyr domain. The cofactor and substrate binding sites of human TKT bear high resemblance to those of other TKTs but also feature unique properties, including two lysines and a serine that interact with the β-phosphate of ThDP. Furthermore, Gln(189) spans over the thiazolium moiety of ThDP and replaces an isoleucine found in most non-mammalian TKTs. The side chain of Gln(428) forms a hydrogen bond with the 4'-amino group of ThDP and replaces a histidine that is invariant in all non-mammalian TKTs. All other amino acids involved in substrate binding and catalysis are strictly conserved. Besides a steady-state kinetic analysis, microscopic equilibria of the donor half-reaction were characterized by an NMR-based intermediate analysis. These studies reveal that formation of the central 1,2-dihydroxyethyl-ThDP carbanion-enamine intermediate is thermodynamically favored with increasing carbon chain length of the donor ketose substrate. Based on the structure of human transketolase and sequence alignments, putative functional properties of the related transketolase-like proteins TKTL1 and -2 are discussed in light of recent findings suggesting that TKTL1 plays a role in cancerogenesis.
Organizational Affiliation:
Institute of Biochemistry and Biotechnology, Martin-Luther-University Halle-Wittenberg, 06120 Halle, Germany.