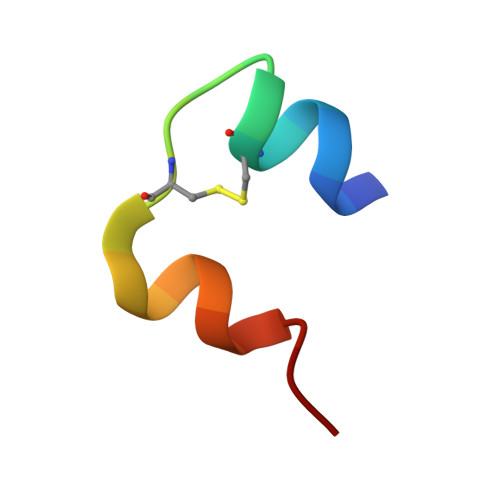
Supramolecular protein engineering: design of zinc-stapled insulin hexamers as a long acting depot.
Phillips, N.B., Wan, Z.L., Whittaker, L., Hu, S.Q., Huang, K., Hua, Q.X., Whittaker, J., Ismail-Beigi, F., Weiss, M.A.(2010) J Biol Chem 285: 11755-11759
- PubMed: 20181952
- DOI: https://doi.org/10.1074/jbc.C110.105825
- Primary Citation of Related Structures:
3KQ6 - PubMed Abstract:
Bottom-up control of supramolecular protein assembly can provide a therapeutic nanobiotechnology. We demonstrate that the pharmacological properties of insulin can be enhanced by design of "zinc staples" between hexamers. Paired (i, i+4) His substitutions were introduced at an alpha-helical surface. The crystal structure contains both classical axial zinc ions and novel zinc ions at hexamer-hexamer interfaces. Although soluble at pH 4, the combined electrostatic effects of the substitutions and bridging zinc ions cause isoelectric precipitation at neutral pH. Following subcutaneous injection in a diabetic rat, the analog effected glycemic control with a time course similar to that of long acting formulation Lantus. Relative to Lantus, however, the analog discriminates at least 30-fold more stringently between the insulin receptor and mitogenic insulin-like growth factor receptor. Because aberrant mitogenic signaling may be associated with elevated cancer risk, such enhanced specificity may improve safety. Zinc stapling provides a general strategy to modify the pharmacokinetic and biological properties of a subcutaneous protein depot.
Organizational Affiliation:
Department of Biochemistry, Western Reserve University School of Medicine, Cleveland, Ohio 44106, USA.