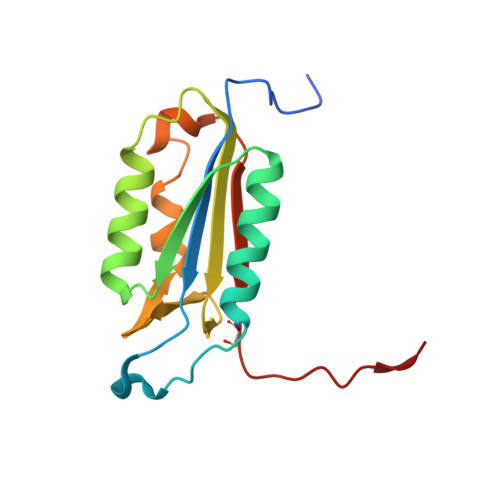
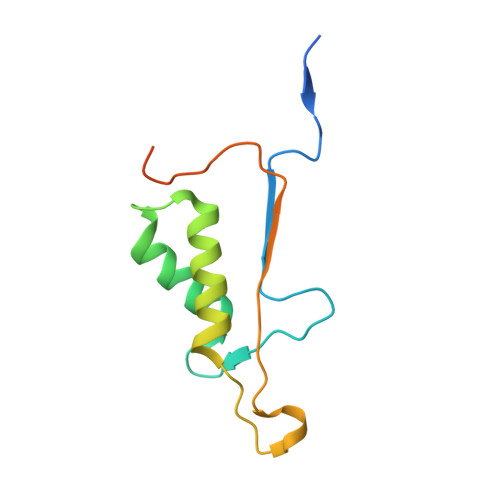
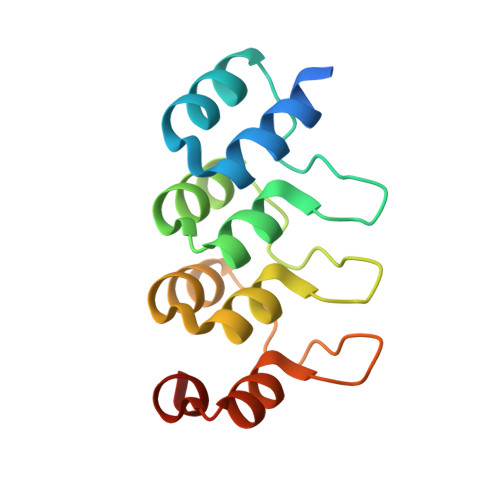
Specific Inhibition of Caspase-3 by a Competitive Darpin: Molecular Mimicry between Native and Designed Inhibitors.
Schroeder, T., Barandun, J., Flutsch, A., Briand, C., Mittl, P., Grutter, M.G.(2013) Structure 21: 277
- PubMed: 23333429
- DOI: https://doi.org/10.1016/j.str.2012.12.011
- Primary Citation of Related Structures:
2XZD, 2Y0B - PubMed Abstract:
Dysregulation of apoptosis is associated with several human diseases. The main apoptotic mediators are caspases, which propagate death signals to downstream targets. Executioner caspase-3 is responsible for the majority of cleavage events and its therapeutic potential is of high interest with to date several available active site peptide inhibitors. These molecules inhibit caspase-3, but also homologous caspases. Here, we describe caspase-3 specific inhibitors D3.4 and D3.8, which have been selected from a library of designed ankyrin repeat proteins (DARPins). The crystal structures of D3.4 and mutants thereof show how high specificity and inhibition is achieved. They also show similarities in the binding mode with that of the natural caspase inhibitor XIAP (X-linked inhibitor of apoptosis). The kinetic data reveal a competitive inhibition mechanism. D3.4 is specific for caspase-3 and does not bind the highly homologous caspase-7. D3.4 therefore is an excellent tool to define the precise role of caspase-3 in the various apoptotic pathways.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland.