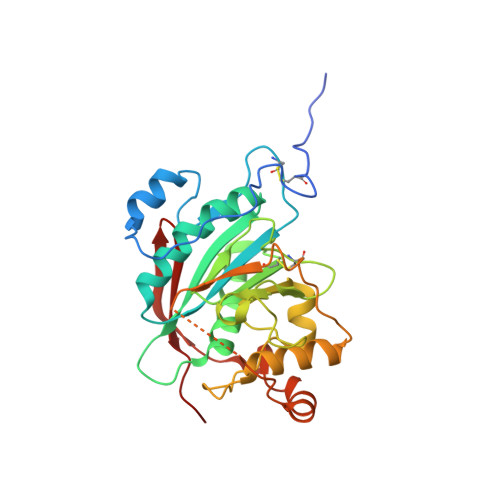
Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
Ramakrishnan, B., Ramasamy, V., Qasba, P.K.(2006) J Mol Biol 357: 1619-1633
- PubMed: 16497331
- DOI: https://doi.org/10.1016/j.jmb.2006.01.088
- Primary Citation of Related Structures:
2FY7, 2FYA, 2FYB, 2FYC, 2FYD - PubMed Abstract:
During the catalytic cycle of beta1,4-galactosyltransferase-1 (Gal-T1), upon the binding of Mn(2+) followed by UDP-Gal, two flexible loops, a long and a short loop, change their conformation from open to closed. We have determined the crystal structures of a human M340H-Gal-T1 mutant in the open conformation (apo-enzyme), its Mn(2+) and Mn(2+)-UDP-Gal-bound complexes, and of a pentenary complex of bovine Gal-T1-Mn(2+)-UDP-GalNAc-Glc-alpha-lactalbumin. These studies show that during the conformational changes in Gal-T1, the coordination of Mn(2+) undergoes significant changes. It loses a coordination bond with a water molecule bound in the open conformation of Gal-T1 while forming a new coordination bond with another water molecule in the closed conformation, creating an active ground-state structure that facilitates enzyme catalysis. In the crystal structure of the pentenary complex, the N-acetylglucosamine (GlcNAc) moiety is found cleaved from UDP-GalNAc and is placed 2.7A away from the O4 oxygen atom of the acceptor Glc molecule, yet to form the product. The anomeric C1 atom of the cleaved GalNAc moiety has only two covalent bonds with its non-hydrogen atoms (O5 and C2 atoms), similar to either an oxocarbenium ion or N-acetylgalactal form, which are crystallographically indistinguishable at the present resolution. The structure also shows that the newly formed, metal-coordinating water molecule forms a hydrogen bond with the beta-phosphate group of the cleaved UDP moiety. This hydrogen bond formation results in the rotation of the beta-phosphate group of UDP away from the cleaved GalNAc moiety, thereby preventing the re-formation of the UDP-sugar during catalysis. Therefore, this water molecule plays an important role during catalysis in ensuring that the catalytic reaction proceeds in a forward direction.
Organizational Affiliation:
Structural Glycobiology Section, Nanobiology Program Center for Cancer Research, NCI-Frederick, Frederick, MD 21702, USA.