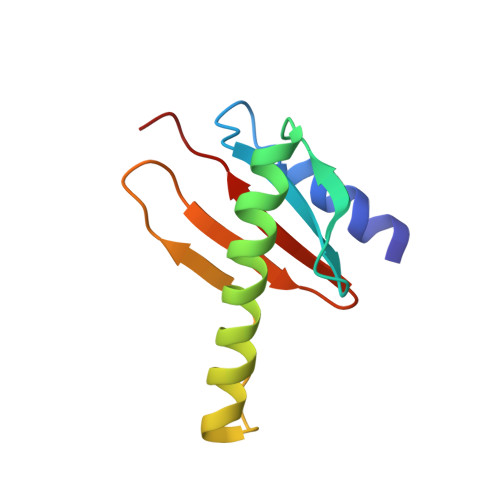
Structure and Dynamics of the Homodimeric Dynein Light Chain km23.
Ilangovan, U., Ding, W., Zhong, Y., Wilson, C.L., Groppe, J.C., Trbovich, J.T., Zuniga, J., Demeler, B., Tang, Q., Gao, G., Mulder, K.M., Hinck, A.P.(2005) J Mol Biol 352 (2): 338-354
- PubMed: 16083906
- DOI: https://doi.org/10.1016/j.jmb.2005.07.002
- Primary Citation of Related Structures:
1Z09 - PubMed Abstract:
km23 (96 residues, 11 kDa) is the mammalian ortholog of Drosophila roadblock, the founding member of LC7/robl/km23 class of dynein light chains. km23 has been shown to be serine-phosphorylated following TGFbeta receptor activation and to bind the dynein intermediate chain in response to such phosphorylation. Here, we report the three-dimensional solution structure of km23, which is shown to be that of a homodimer, similar to that observed for the heterodimeric complex formed between p14 and MP1, two distantly related members of the MglB/robl superfamily, but distinct from the LC8 and Tctex-1 classes of dynein light chains, which also adopt homodimeric structures. The conserved surface residues of km23, including three serine residues, are located predominantly on a single face of the molecule. Adjacent to this face is a large cleft formed by the incomplete overlap of loops from opposite monomers. As shown by NMR relaxation data collected at two fields, several cleft residues are flexible on the ns-ps and ms-mus timescales. Based on these observations, we propose that the patch of conserved residues on the central face of the molecule corresponds to the site at which km23 binds the dynein intermediate chain and that the flexible cleft formed between the overlap of loops from the two monomers corresponds to the site at which km23 binds other partners, such as the TGFbeta type II receptor or Smad2.
Organizational Affiliation:
Department of Biochemistry, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, USA.