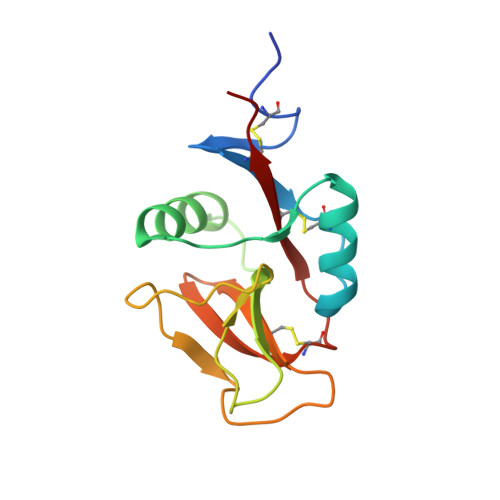
The 1.4 angstrom crystal structure of the human oxidized low density lipoprotein receptor lox-1.
Park, H., Adsit, F.G., Boyington, J.C.(2005) J Biol Chem 280: 13593-13599
- PubMed: 15695803
- DOI: https://doi.org/10.1074/jbc.M500768200
- Primary Citation of Related Structures:
1YPO, 1YPQ, 1YPU - PubMed Abstract:
The lectin-like oxidized low density lipoprotein receptor-1 (Lox-1) mediates the recognition and internalization of oxidatively modified low density lipoprotein by vascular endothelial cells. This interaction results in a number of pro-atherogenic cellular responses that probably play a significant role in the pathology of atherosclerosis. The 1.4 angstrom crystal structure of the extracellular C-type lectin-like domain of human Lox-1 reveals a heart-shaped homodimer with a ridge of six basic amino acids extending diagonally across the apolar top of Lox-1, a central hydrophobic tunnel that extends through the entire molecule, and an electrostatically neutral patch of 12 charged residues that resides next to the tunnel at each opening. Based on the arrangement of critical binding residues on the Lox-1 structure, we propose a binding mode for the recognition of modified low density lipoprotein and other Lox-1 ligands.
Organizational Affiliation:
Biomolecular Crystallography Group, Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, North Carolina 27709, USA.