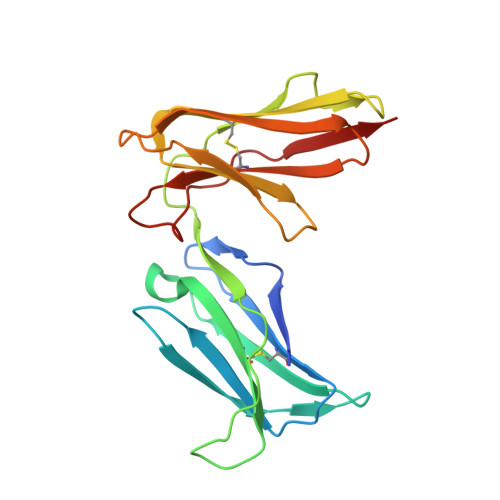
Crystal structure of the human natural killer cell activating receptor KIR2DS2 (CD158j)
Saulquin, X., Gastinel, L.N., Vivier, E.(2003) J Exp Med 197: 933-938
- PubMed: 12668644
- DOI: https://doi.org/10.1084/jem.20021624
- Primary Citation of Related Structures:
1M4K - PubMed Abstract:
Killer cell Ig-like receptors (KIRs) regulate the function of human natural killer and T cell subsets. A feature of the KIR locus is the clustering of homologous genes encoding for inhibitory and activating KIR. Inhibitory and activating KIR differ for ligand specificities and/or affinities. In particular, we show here with KIR tetramers that activating KIR2DS2 does not bind HLA-Cw3 molecules recognized by inhibitory KIR2DL2, despite 99% extracellular amino acid identity. We also report the 2.3-A structure of KIR2DS2, which reveals subtle displacements of two residues (Tyr45 and Gln71) involved in the interaction of KIR2DL2 with HLA-Cw3. These results show that KIR molecules cannot tolerate any variability in their three-dimensional structure without altering their MHC class I recognition capacities. Therefore, the mode of recognition used by KIR largely differs from the conformational changes that characterize T cell receptor or NKG2D interaction with their respective ligands.
Organizational Affiliation:
Centre d'Immunologie INSERM-CNRS, de Marseille Luminy, Case 906, Parc Scientifique de Luminy, 13288 Marseille Cedex 09, France.