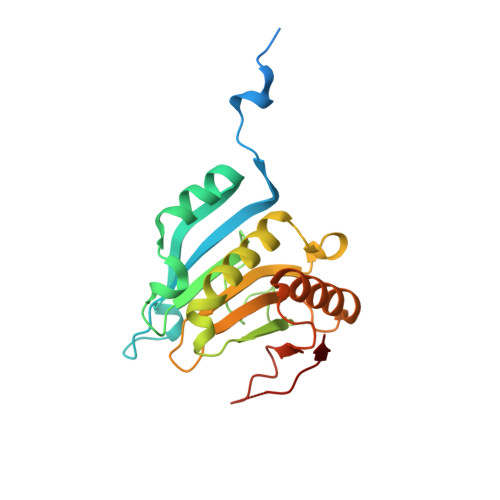
Crystal structures of 7-methylguanosine 5'-triphosphate (m(7)GTP)- and P(1)-7-methylguanosine-P(3)-adenosine-5',5'-triphosphate (m(7)GpppA)-bound human full-length eukaryotic initiation factor 4E: biological importance of the C-terminal flexible region
Tomoo, K., Shen, X., Okabe, K., Nozoe, Y., Fukuhara, S., Morino, S., Ishida, T., Taniguchi, T., Hasegawa, H., Terashima, A., Sasaki, M., Katsuya, Y., Kitamura, K., Miyoshi, H., Ishikawa, M., Miura, K.(2002) Biochem J 362: 539-544
- PubMed: 11879179
- DOI: https://doi.org/10.1042/0264-6021:3620539
- Primary Citation of Related Structures:
1IPB, 1IPC - PubMed Abstract:
The crystal structures of the full-length human eukaryotic initiation factor (eIF) 4E complexed with two mRNA cap analogues [7-methylguanosine 5'-triphosphate (m(7)GTP) and P(1)-7-methylguanosine-P(3)-adenosine-5',5'-triphosphate (m(7)GpppA)] were determined at 2.0 A resolution (where 1 A=0.1 nm). The flexibility of the C-terminal loop region of eIF4E complexed with m(7)GTP was significantly reduced when complexed with m(7)GpppA, suggesting the importance of the second nucleotide in the mRNA cap structure for the biological function of eIF4E, especially the fixation and orientation of the C-terminal loop region, including the eIF4E phosphorylation residue. The present results provide the structural basis for the biological function of both N- and C-terminal mobile regions of eIF4E in translation initiation, especially the regulatory function through the switch-on/off of eIF4E-binding protein-eIF4E phosphorylation.
Organizational Affiliation:
Department of Physical Chemistry, Osaka University of Pharmaceutical Sciences, 4-20-1 Nasahara, Takatsuki, Osaka 569-1094, Japan. tomoo@gly.oups.ac.jp