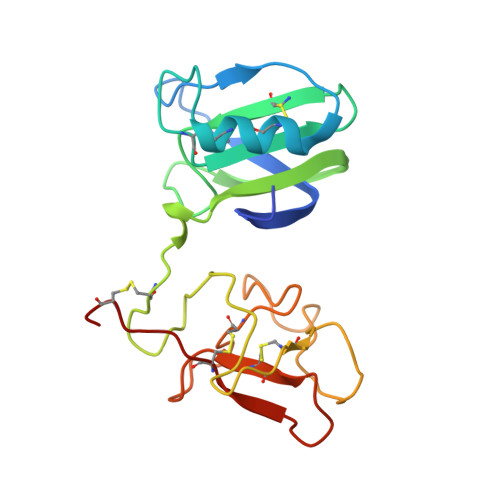
A New Crystal Form of the Nk1 Splice Variant of Hgf/Sf Demonstrates Extensive Hinge Movement and Suggests that the Nk1 Dimer Originates by Domain Swapping
Watanabe, K., Chirgadze, D.Y., Lietha, D., De Jonge, H., Blundell, T.L., Gherardi, E.(2002) J Mol Biol 319: 283
- PubMed: 12051906
- DOI: https://doi.org/10.1016/S0022-2836(02)00199-7
- Primary Citation of Related Structures:
1GP9 - PubMed Abstract:
NK1 is a splice variant of the polypeptide growth factor HGF/SF that consists of the N terminal (N) and first kringle (K) domains and retains receptor binding and signalling. While NK1 behaves as a monomer in solution, two independent crystallographic structures have previously shown an identical, tightly packed dimer. Here we report a novel orthorhombic crystal form of NK1 at 2.5 A resolution in which four NK1 protomers are packed in two distinct dimers in the asymmetric unit. Although the basic architecture of the new NK1 dimers is similar to the two described earlier, the new crystal form demonstrates extensive hinge movement between the N and K domain that leads to re-orientation of the receptor-binding sites. The hinge bending is evidence of the paucity of strong interactions between domains within the protomer, in contrast to the extensive interactions between protomers in the dimer. These observations are consistent with domain swapping in the dimer, such that the interdomain interactions of the monomer are replaced by equivalent interprotomer interactions in the dimer and offer a route for protein engineering of NK1 variants which may act as receptor antagonists.
Organizational Affiliation:
Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, UK.