8E57
Rabbit L-type voltage-gated calcium channel Cav1.1 in the presence of Amiodarone and 100 microM MNI-1 at 2.8 Angstrom resolution
- PDB DOI: https://doi.org/10.2210/pdb8E57/pdb
- EM Map EMD-27905: EMDB EMDataResource
- Classification: TRANSPORT PROTEIN
- Organism(s): Oryctolagus cuniculus
- Mutation(s): No
- Membrane Protein: Yes OPMPDBTMmpstruc
- Deposited: 2022-08-20 Released: 2022-12-07
- Funding Organization(s): National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS)
Experimental Data Snapshot
- Method: ELECTRON MICROSCOPY
- Resolution: 2.80 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
wwPDB Validation 3D Report Full Report
This is version 1.1 of the entry. See complete history.
Macromolecules
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Voltage-dependent L-type calcium channel subunit alpha-1S | 1,873 | Oryctolagus cuniculus | Mutation(s): 0 Membrane Entity: Yes | 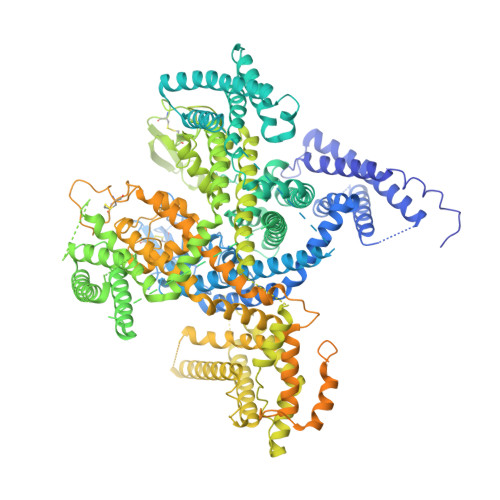 | |
UniProt | |||||
Find proteins for P07293 (Oryctolagus cuniculus) Explore P07293 Go to UniProtKB: P07293 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P07293 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Voltage-dependent calcium channel gamma-1 subunit | B [auth E] | 222 | Oryctolagus cuniculus | Mutation(s): 0 Membrane Entity: Yes | 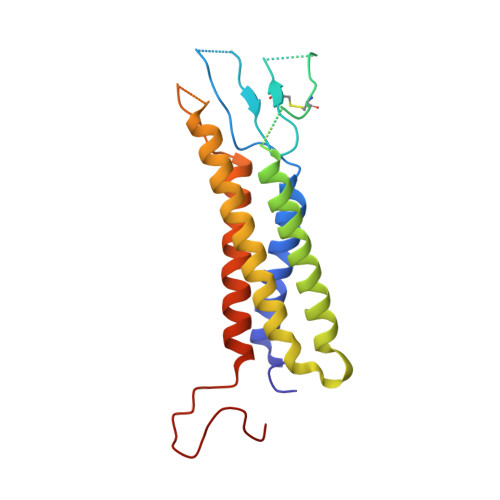 |
UniProt | |||||
Find proteins for P19518 (Oryctolagus cuniculus) Explore P19518 Go to UniProtKB: P19518 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P19518 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Voltage-dependent calcium channel subunit alpha-2/delta-1 | C [auth F] | 1,106 | Oryctolagus cuniculus | Mutation(s): 0 Membrane Entity: Yes | 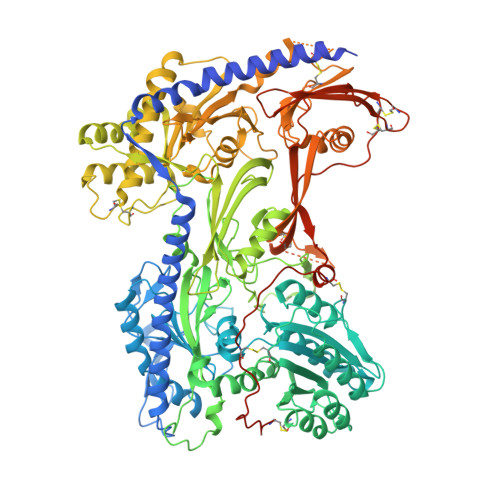 |
UniProt | |||||
Find proteins for P13806 (Oryctolagus cuniculus) Explore P13806 Go to UniProtKB: P13806 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P13806 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Oligosaccharides
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | D [auth B], E [auth C], F [auth D], G | 2 | N/A | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | H | 3 | N/A | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G47362BJ GlyCosmos: G47362BJ GlyGen: G47362BJ | |||||
Small Molecules
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BBI (Subject of Investigation/LOI) Query on BBI | L [auth A] | (2-butyl-1-benzofuran-3-yl){4-[2-(diethylamino)ethoxy]-3,5-diiodophenyl}methanone C25 H29 I2 N O3 IYIKLHRQXLHMJQ-UHFFFAOYSA-N |  | ||
| WFR (Subject of Investigation/LOI) Query on WFR | M [auth A] | propan-2-yl (2S)-2-{[(S)-{[(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-4-ethynyl-3-hydroxy-4-methyloxolan-2-yl]methoxy}(phenoxy)phosphoryl]amino}propanoate (non-preferred name) C24 H30 N3 O9 P HVOMZYITEDRMBB-XSZHHMMYSA-N |  | ||
| NAG Query on NAG | O [auth F] P [auth F] Q [auth F] R [auth F] S [auth F] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| CA Query on CA | K [auth A], N [auth F] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
Biologically Interesting Molecules (External Reference) 1 Unique
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900017 Query on PRD_900017 | H | triacetyl-beta-chitotriose | Oligosaccharide / Inhibitor |  | |
Experimental Data & Validation
Experimental Data
- Method: ELECTRON MICROSCOPY
- Resolution: 2.80 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
Entry History & Funding Information
Deposition Data
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | 5R01GM130762 |
Revision History (Full details and data files)
- Version 1.0: 2022-12-07
Type: Initial release - Version 1.1: 2022-12-21
Changes: Database references













