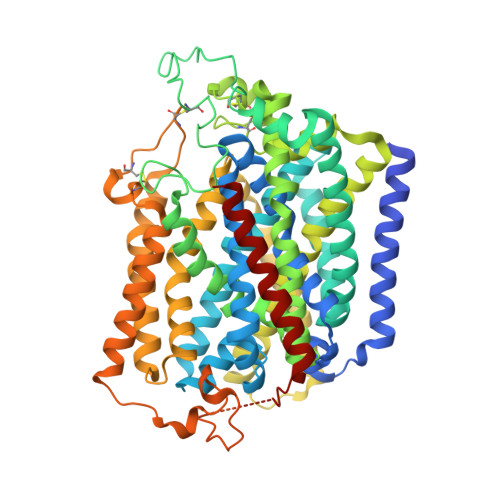
Structural basis of inhibition of the human SGLT2-MAP17 glucose transporter.
Niu, Y., Liu, R., Guan, C., Zhang, Y., Chen, Z., Hoerer, S., Nar, H., Chen, L.(2022) Nature 601: 280-284
- PubMed: 34880493
- DOI: https://doi.org/10.1038/s41586-021-04212-9
- Primary Citation of Related Structures:
7VSI - PubMed Abstract:
Human sodium-glucose cotransporter 2 (hSGLT2) mediates the reabsorption of the majority of filtrated glucose in the kidney 1 . Pharmacological inhibition of hSGLT2 by oral small-molecule inhibitors, such as empagliflozin, leads to enhanced excretion of glucose and is widely used in the clinic to manage blood glucose levels for the treatment of type 2 diabetes 1 . Here we determined the cryogenic electron microscopy structure of the hSGLT2-MAP17 complex in the empagliflozin-bound state to an overall resolution of 2.95 Å. Our structure shows eukaryotic SGLT-specific structural features. MAP17 interacts with transmembrane helix 13 of hSGLT2. Empagliflozin occupies both the sugar-substrate-binding site and the external vestibule to lock hSGLT2 in an outward-open conformation, thus inhibiting the transport cycle. Our work provides a framework for understanding the mechanism of SLC5A family glucose transporters and also develops a foundation for the future rational design and optimization of new inhibitors targeting these transporters.
Organizational Affiliation:
State Key Laboratory of Membrane Biology, College of Future Technology, Institute of Molecular Medicine, Peking University, Beijing Key Laboratory of Cardiometabolic Molecular Medicine, Beijing, China.