6Y1Z
Mouse serotonin 5HT3 receptor in complex with palonosetron
- PDB DOI: https://doi.org/10.2210/pdb6Y1Z/pdb
- EM Map EMD-10673: EMDB EMDataResource
- Classification: TRANSPORT PROTEIN
- Organism(s): Mus musculus
- Expression System: Homo sapiens
- Mutation(s): No
- Membrane Protein: Yes OPMPDBTMMemProtMDmpstruc
- Deposited: 2020-02-14 Released: 2020-03-04
- Funding Organization(s): European Research Council (ERC)
Experimental Data Snapshot
- Method: ELECTRON MICROSCOPY
- Resolution: 2.82 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
wwPDB Validation 3D Report Full Report
This is version 3.0 of the entry. See complete history.
Macromolecules
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 5-hydroxytryptamine receptor 3A | A, B [auth D], C [auth B], D [auth C], E | 566 | Mus musculus | Mutation(s): 0 Gene Names: Htr3a, 5ht3, Htr3 Membrane Entity: Yes | 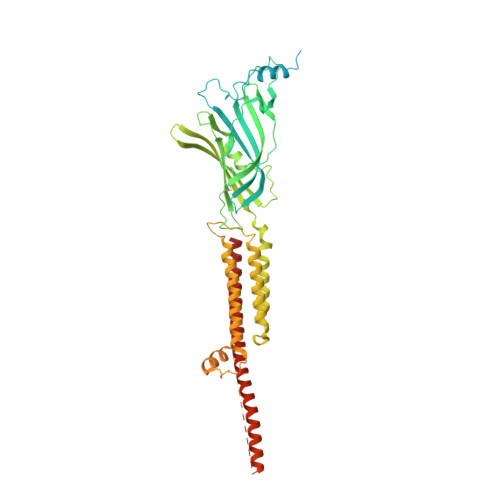 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P23979 (Mus musculus) Explore P23979 Go to UniProtKB: P23979 | |||||
IMPC: MGI:96282 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P23979 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Oligosaccharides
Small Molecules
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| O7B (Subject of Investigation/LOI) Query on O7B | AA [auth B], BA [auth C], FA [auth E], S [auth A], W [auth D] | (3~{a}~{S})-2-[(3~{S})-1-azabicyclo[2.2.2]octan-3-yl]-3~{a},4,5,6-tetrahydro-3~{H}-benzo[de]isoquinolin-1-one C19 H24 N2 O CPZBLNMUGSZIPR-NVXWUHKLSA-N |  | ||
| NAG Query on NAG | EA [auth C], IA [auth E], R [auth A], V [auth D], Z [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| TRP Query on TRP | CA [auth C], GA [auth E], P [auth A], T [auth D], X [auth B] | TRYPTOPHAN C11 H12 N2 O2 QIVBCDIJIAJPQS-VIFPVBQESA-N |  | ||
| HIS Query on HIS | DA [auth C], HA [auth E], Q [auth A], U [auth D], Y [auth B] | HISTIDINE C6 H10 N3 O2 HNDVDQJCIGZPNO-YFKPBYRVSA-O |  | ||
Experimental Data & Validation
Experimental Data
- Method: ELECTRON MICROSCOPY
- Resolution: 2.82 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC | v2.13 |
| MODEL REFINEMENT | PHENIX | 1.17 |
Entry History & Funding Information
Deposition Data
- Released Date: 2020-03-04 Deposition Author(s): Zarkadas, E., Perot, J., Nury, H.
| Funding Organization | Location | Grant Number |
|---|---|---|
| European Research Council (ERC) | France | 637733 |
Revision History (Full details and data files)
- Version 1.0: 2020-03-04
Type: Initial release - Version 2.0: 2020-07-29
Type: Remediation
Reason: Carbohydrate remediation
Changes: Advisory, Atomic model, Data collection, Derived calculations, Structure summary - Version 2.1: 2020-08-12
Changes: Database references, Structure summary - Version 2.2: 2020-10-14
Changes: Database references - Version 3.0: 2022-09-14
Changes: Advisory, Atomic model, Data collection, Database references, Derived calculations, Polymer sequence, Source and taxonomy, Structure summary















