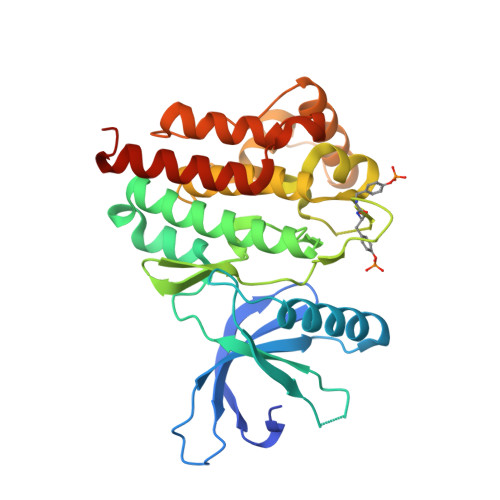
Degradation of Janus kinases in CRLF2-rearranged acute lymphoblastic leukemia.
Chang, Y., Min, J., Jarusiewicz, J.A., Actis, M., Yu-Chen Bradford, S., Mayasundari, A., Yang, L., Chepyala, D., Alcock, L.J., Roberts, K.G., Nithianantham, S., Maxwell, D., Rowland, L., Larsen, R., Seth, A., Goto, H., Imamura, T., Akahane, K., Hansen, B.S., Pruett-Miller, S.M., Paietta, E.M., Litzow, M.R., Qu, C., Yang, J.J., Fischer, M., Rankovic, Z., Mullighan, C.G.(2021) Blood 138: 2313-2326
- PubMed: 34110416
- DOI: https://doi.org/10.1182/blood.2020006846
- Primary Citation of Related Structures:
6WTN, 6WTO, 6WTP, 6WTQ - PubMed Abstract:
CRLF2-rearranged (CRLF2r) acute lymphoblastic leukemia (ALL) accounts for more than half of Philadelphia chromosome-like (Ph-like) ALL and is associated with a poor outcome in children and adults. Overexpression of CRLF2 results in activation of Janus kinase (JAK)-STAT and parallel signaling pathways in experimental models, but existing small molecule inhibitors of JAKs show variable and limited efficacy. Here, we evaluated the efficacy of proteolysis-targeting chimeras (PROTACs) directed against JAKs. Solving the structure of type I JAK inhibitors ruxolitinib and baricitinib bound to the JAK2 tyrosine kinase domain enabled the rational design and optimization of a series of cereblon (CRBN)-directed JAK PROTACs utilizing derivatives of JAK inhibitors, linkers, and CRBN-specific molecular glues. The resulting JAK PROTACs were evaluated for target degradation, and activity was tested in a panel of leukemia/lymphoma cell lines and xenograft models of kinase-driven ALL. Multiple PROTACs were developed that degraded JAKs and potently killed CRLF2r cell lines, the most active of which also degraded the known CRBN neosubstrate GSPT1 and suppressed proliferation of CRLF2r ALL in vivo, e.g. compound 7 (SJ988497). Although dual JAK/GSPT1-degrading PROTACs were the most potent, the development and evaluation of multiple PROTACs in an extended panel of xenografts identified a potent JAK2-degrading, GSPT1-sparing PROTAC that demonstrated efficacy in the majority of kinase-driven xenografts that were otherwise unresponsive to type I JAK inhibitors, e.g. compound 8 (SJ1008030). Together, these data show the potential of JAK-directed protein degradation as a therapeutic approach in JAK-STAT-driven ALL and highlight the interplay of JAK and GSPT1 degradation activity in this context.
Organizational Affiliation:
Department of Pathology.