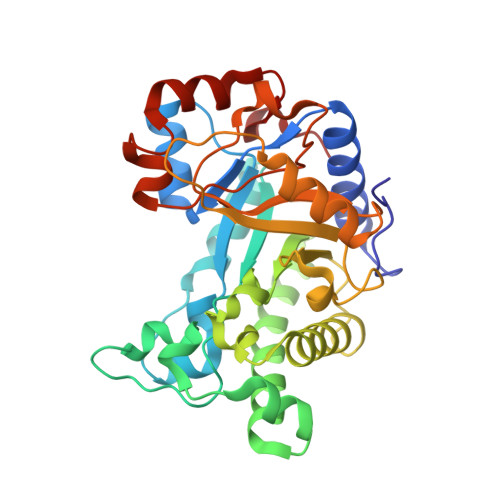
Crystal structure and substrate interactions of an unusual fungal non-CBM carrying GH26 endo-beta-mannanase from Yunnania penicillata.
von Freiesleben, P., Moroz, O.V., Blagova, E., Wiemann, M., Spodsberg, N., Agger, J.W., Davies, G.J., Wilson, K.S., Stalbrand, H., Meyer, A.S., Krogh, K.B.R.M.(2019) Sci Rep 9: 2266-2266
- PubMed: 30783168
- DOI: https://doi.org/10.1038/s41598-019-38602-x
- Primary Citation of Related Structures:
6HPF - PubMed Abstract:
Endo-β(1 → 4)-mannanases (endomannanases) catalyse degradation of β-mannans, an abundant class of plant polysaccharides. This study investigates structural features and substrate binding of YpenMan26A, a non-CBM carrying endomannanase from Yunnania penicillata. Structural and sequence comparisons to other fungal family GH26 endomannanases showed high sequence similarities and conserved binding residues, indicating that fungal GH26 endomannanases accommodate galactopyranosyl units in the -3 and -2 subsites. Two striking amino acid differences in the active site were found when the YpenMan26A structure was compared to a homology model of Wsp.Man26A from Westerdykella sp. and the sequences of nine other fungal GH26 endomannanases. Two YpenMan26A mutants, W110H and D37T, inspired by differences observed in Wsp.Man26A, produced a shift in how mannopentaose bound across the active site cleft and a decreased affinity for galactose in the -2 subsite, respectively, compared to YpenMan26A. YpenMan26A was moreover found to have a flexible surface loop in the position where PansMan26A from Podospora anserina has an α-helix (α9) which interacts with its family 35 CBM. Sequence alignment inferred that the core structure of fungal GH26 endomannanases differ depending on the natural presence of this type of CBM. These new findings have implications for selecting and optimising these enzymes for galactomannandegradation.
Organizational Affiliation:
Novozymes A/S, Krogshøjvej 36, 2880, Bagsværd, Denmark.