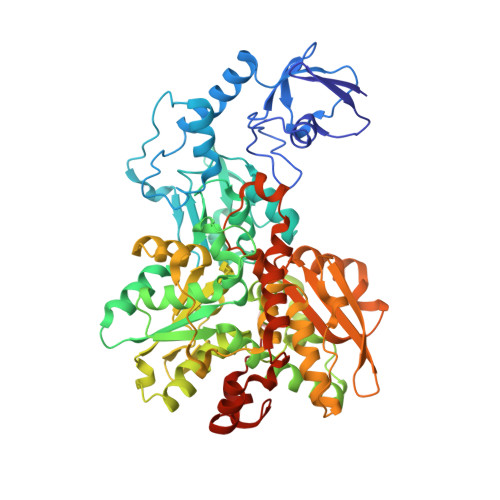
Crystallographic and spectroscopic assignment of the proton transfer pathway in [FeFe]-hydrogenases.
Duan, J., Senger, M., Esselborn, J., Engelbrecht, V., Wittkamp, F., Apfel, U.P., Hofmann, E., Stripp, S.T., Happe, T., Winkler, M.(2018) Nat Commun 9: 4726-4726
- PubMed: 30413719
- DOI: https://doi.org/10.1038/s41467-018-07140-x
- Primary Citation of Related Structures:
6GLY, 6GLZ, 6GM0, 6GM1, 6GM2, 6GM3, 6GM4, 6GM5, 6GM6, 6GM7, 6GM8 - PubMed Abstract:
The unmatched catalytic turnover rates of [FeFe]-hydrogenases require an exceptionally efficient proton-transfer (PT) pathway to shuttle protons as substrates or products between bulk water and catalytic center. For clostridial [FeFe]-hydrogenase CpI such a pathway has been proposed and analyzed, but mainly on a theoretical basis. Here, eleven enzyme variants of two different [FeFe]-hydrogenases (CpI and HydA1) with substitutions in the presumptive PT-pathway are examined kinetically, spectroscopically, and crystallographically to provide solid experimental proof for its role in hydrogen-turnover. Targeting key residues of the PT-pathway by site directed mutagenesis significantly alters the pH-activity profile of these variants and in presence of H 2 their cofactor is trapped in an intermediate state indicative of precluded proton-transfer. Furthermore, crystal structures coherently explain the individual levels of residual activity, demonstrating e.g. how trapped H 2 O molecules rescue the interrupted PT-pathway. These features provide conclusive evidence that the targeted positions are indeed vital for catalytic proton-transfer.
Organizational Affiliation:
Department of Plant Biochemistry, Photobiotechnology, Ruhr-Universität Bochum, 44801, Bochum, Germany.