Crystal Structure of Human PPARgamma Ligand Binding Domain in Complex with TRAP220 Coactivator Peptide
Shang, J., Kojetin, D.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peroxisome proliferator-activated receptor gamma | 297 | Homo sapiens | Mutation(s): 0 Gene Names: PPARG, NR1C3 | 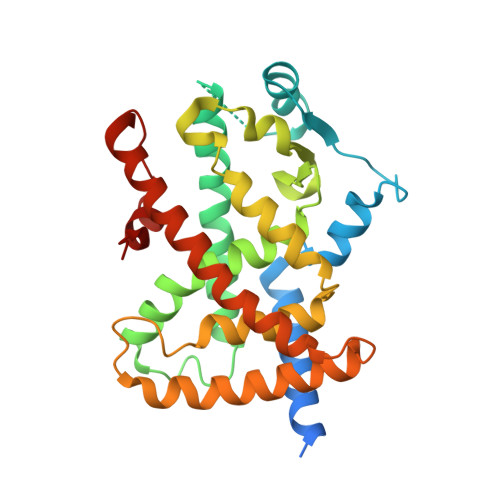 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P37231 (Homo sapiens) Explore P37231 Go to UniProtKB: P37231 | |||||
PHAROS: P37231 GTEx: ENSG00000132170 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P37231 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| TRAP220 Coactivator Peptide (Mediator of RNA polymerase II transcription subunit 1) | 19 | Homo sapiens | Mutation(s): 0 Gene Names: MED1 |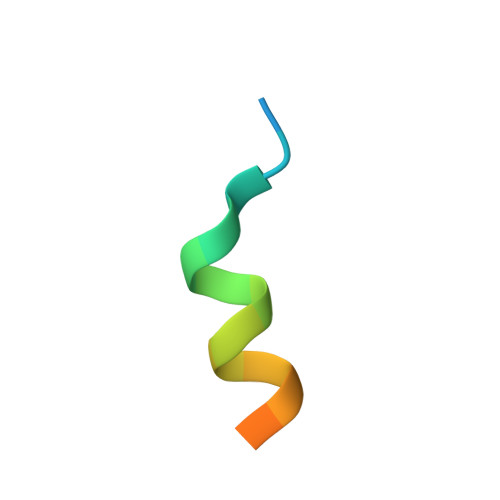  | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 88.09 | α = 90 |
| b = 54.73 | β = 90.05 |
| c = 123.471 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| MOSFLM | data reduction |
| SCALA | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of Diabetes and Digestive and Kidney Disease (NIH/NIDDK) | United States | R01DK101871 |