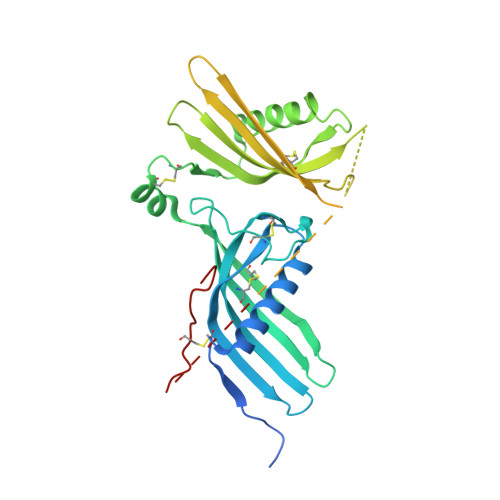
The C-terminal region of human plasma fetuin-B is dispensable for the raised-elephant-trunk mechanism of inhibition of astacin metallopeptidases.
Guevara, T., Korschgen, H., Cuppari, A., Schmitz, C., Kuske, M., Yiallouros, I., Floehr, J., Jahnen-Dechent, W., Stocker, W., Gomis-Ruth, F.X.(2019) Sci Rep 9: 14683-14683
- PubMed: 31604990
- DOI: https://doi.org/10.1038/s41598-019-51095-y
- Primary Citation of Related Structures:
6SAZ - PubMed Abstract:
Human fetuin-B plays a key physiological role in human fertility through its inhibitory action on ovastacin, a member of the astacin family of metallopeptidases. The inhibitor consists of tandem cystatin-like domains (CY1 and CY2), which are connected by a linker containing a "CPDCP-trunk" and followed by a C-terminal region (CTR) void of regular secondary structure. Here, we solved the crystal structure of the complex of the inhibitor with archetypal astacin from crayfish, which is a useful model of human ovastacin. Two hairpins from CY2, the linker, and the tip of the "legumain-binding loop" of CY1 inhibit crayfish astacin following the "raised-elephant-trunk mechanism" recently reported for mouse fetuin-B. This inhibition is exerted by blocking active-site cleft sub-sites upstream and downstream of the catalytic zinc ion, but not those flanking the scissile bond. However, contrary to the mouse complex, which was obtained with fetuin-B nicked at a single site but otherwise intact, most of the CTR was proteolytically removed during crystallization of the human complex. Moreover, the two complexes present in the crystallographic asymmetric unit diverged in the relative arrangement of CY1 and CY2, while the two complexes found for the mouse complex crystal structure were equivalent. Biochemical studies in vitro confirmed the differential cleavage susceptibility of human and mouse fetuin-B in front of crayfish astacin and revealed that the cleaved human inhibitor blocks crayfish astacin and human meprin α and β only slightly less potently than the intact variant. Therefore, the CTR of animal fetuin-B orthologs may have a function in maintaining a particular relative orientation of CY1 and CY2 that nonetheless is dispensable for peptidase inhibition.
Organizational Affiliation:
Proteolysis Lab, Department of Structural Biology, Molecular Biology Institute of Barcelona, CSIC, Barcelona Science Park, Helix Building, c/ Baldiri Reixac, 15-21, E-08028, Barcelona, Catalonia, Spain.