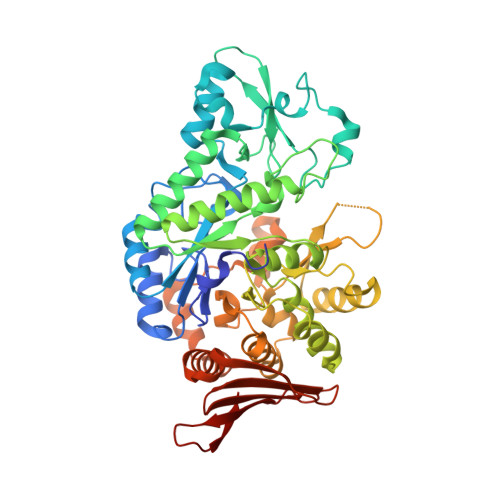
Structural Comparison of a Promiscuous and a Highly Specific Sucrose 6 F -Phosphate Phosphorylase.
Franceus, J., Capra, N., Desmet, T., Thunnissen, A.W.H.(2019) Int J Mol Sci 20
- PubMed: 31405215
- DOI: https://doi.org/10.3390/ijms20163906
- Primary Citation of Related Structures:
6S9U, 6S9V - PubMed Abstract:
In family GH13 of the carbohydrate-active enzyme database, subfamily 18 contains glycoside phosphorylases that act on α-sugars and glucosides. Because their phosphorolysis reactions are effectively reversible, these enzymes are of interest for the biocatalytic synthesis of various glycosidic compounds. Sucrose 6 F -phosphate phosphorylases (SPPs) constitute one of the known substrate specificities. Here, we report the characterization of an SPP from Ilumatobacter coccineus with a far stricter specificity than the previously described promiscuous SPP from Thermoanaerobacterium thermosaccharolyticum . Crystal structures of both SPPs were determined to provide insight into their similarities and differences. The residues responsible for binding the fructose 6-phosphate group in subsite +1 were found to differ considerably between the two enzymes. Furthermore, several variants that introduce a higher degree of substrate promiscuity in the strict SPP from I. coccineus were designed. These results contribute to an expanded structural knowledge of enzymes in subfamily GH13_18 and facilitate their rational engineering.
Organizational Affiliation:
Centre for Synthetic Biology (CSB), Department of Biotechnology, Ghent University, Coupure Links 653, 9000 Ghent, Belgium. jorick.franceus@ugent.be.