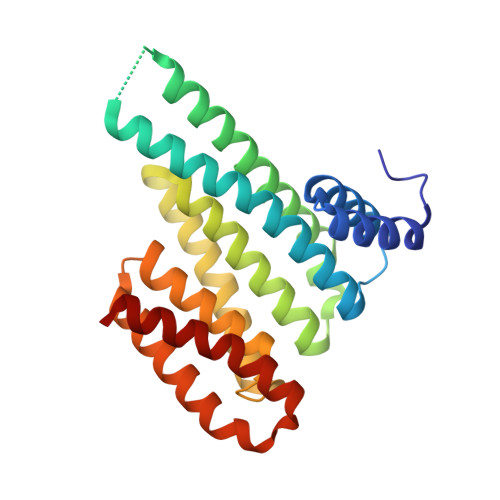
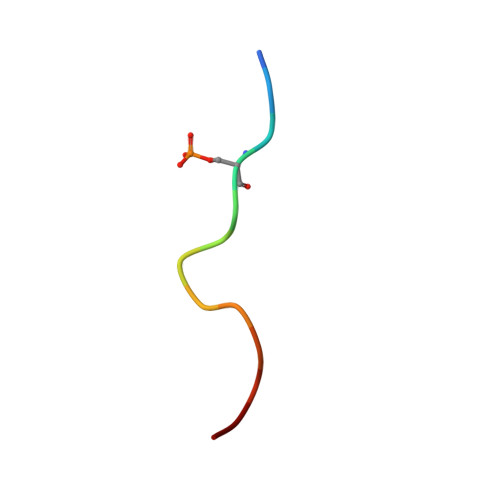Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
Guillory, X., Wolter, M., Leysen, S., Neves, J.F., Kuusk, A., Genet, S., Somsen, B., Morrow, J.K., Rivers, E., van Beek, L., Patel, J., Goodnow, R., Schoenherr, H., Fuller, N., Cao, Q., Doveston, R.G., Brunsveld, L., Arkin, M.R., Castaldi, P., Boyd, H., Landrieu, I., Chen, H., Ottmann, C.(2020) J Med Chem 63: 6694-6707
- PubMed: 32501690
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01942
- Primary Citation of Related Structures:
6R5L, 6RHC, 6RJL, 6RJQ, 6RJZ, 6RK8, 6RKI, 6RKK, 6RKM, 6RL3, 6RL4, 6RL6, 6RM5, 6RM7, 6RP6, 6RWH, 6RWI, 6RWS, 6RWU, 6RX2, 6S39, 6S3C, 6S40, 6S9Q, 6SIN, 6SIO, 6SIP, 6SIQ, 6SLV, 6SLW, 6SLX - PubMed Abstract:
Stabilization of protein-protein interactions (PPIs) holds great potential for therapeutic agents, as illustrated by the successful drugs rapamycin and lenalidomide. However, how such interface-binding molecules can be created in a rational, bottom-up manner is a largely unanswered question. We report here how a fragment-based approach can be used to identify chemical starting points for the development of small-molecule stabilizers that differentiate between two different PPI interfaces of the adapter protein 14-3-3. The fragments discriminately bind to the interface of 14-3-3 with the recognition motif of either the tumor suppressor protein p53 or the oncogenic transcription factor TAZ. This X-ray crystallography driven study shows that the rim of the interface of individual 14-3-3 complexes can be targeted in a differential manner with fragments that represent promising starting points for the development of specific 14-3-3 PPI stabilizers.
Organizational Affiliation:
Laboratory of Chemical Biology, Department of Biomedical Engineering and Institute for Complex Molecular Systems, Eindhoven University of Technology, P.O. Box 513, 5600MB Eindhoven, The Netherlands.