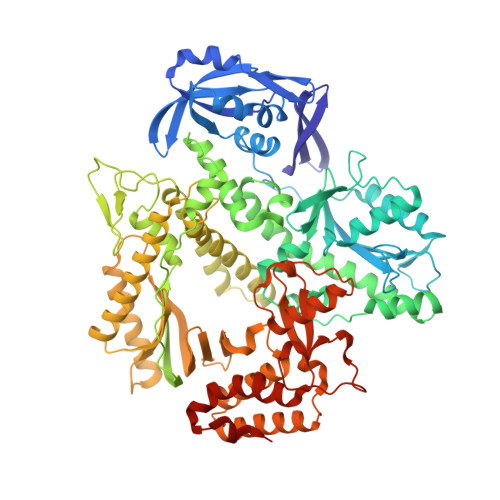
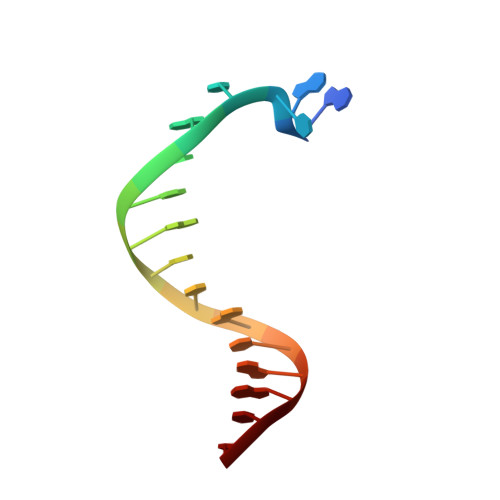
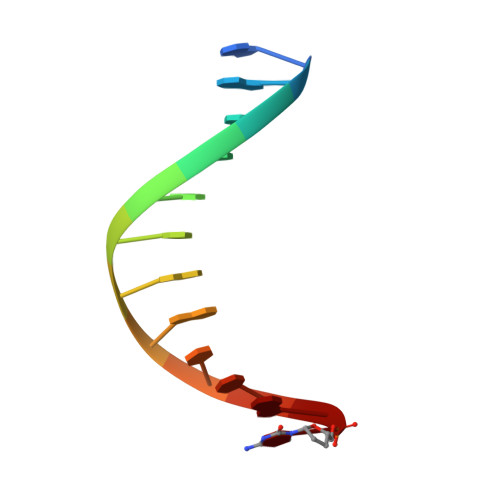
The Structure of an Archaeal B-Family DNA Polymerase in Complex with a Chemically Modified Nucleotide.
Kropp, H.M., Diederichs, K., Marx, A.(2019) Angew Chem Int Ed Engl 58: 5457-5461
- PubMed: 30761722
- DOI: https://doi.org/10.1002/anie.201900315
- Primary Citation of Related Structures:
6Q4T, 6Q4U, 6Q4V - PubMed Abstract:
Archaeal B-family DNA polymerases (DNA pols) are the driving force of cutting-edge biotechnological applications like next-generation sequencing. The acceptance of chemically modified nucleotides by DNA pols is key to these technologies. Until now, no structural data have been available for these DNA pols in complex with modified substrates, which could build the basis for understanding interactions between the enzyme and the chemically modified nucleotide and for the further development of next-generation nucleotides. For the first time, we crystallized an exonuclease-deficient variant of the wild-type B-family KOD DNA pol with a modified nucleotide in a closed, ternary complex. We also crystalized the A-family DNA pol KlenTaq with the same nucleotide. The reported structural data reveal how the protein and the DNA modulate two distinct conformations of the appended moiety in the A- and B-family DNA pols and how these influence the processing of the modified nucleotide. Overall, this study provides first insight into the interplay between B-family DNA pols and relevant modified substrates.
Organizational Affiliation:
Department of Chemistry and Konstanz Research School Chemical Biology, University of Konstanz, Universitätsstrasse 10, 7857, Konstanz, Germany.