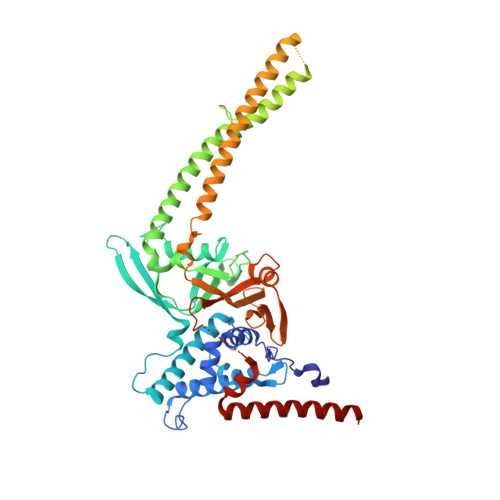
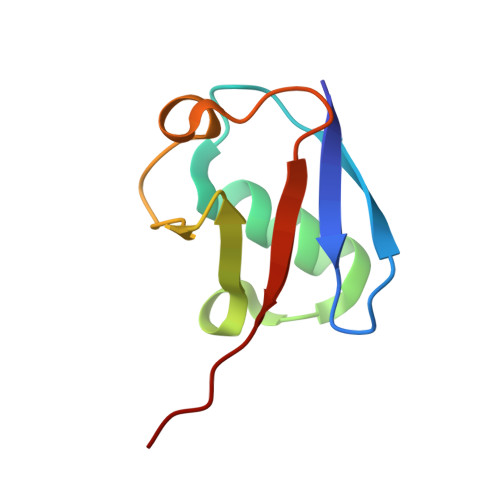
Differential Oligomerization of the Deubiquitinases USP25 and USP28 Regulates Their Activities.
Sauer, F., Klemm, T., Kollampally, R.B., Tessmer, I., Nair, R.K., Popov, N., Kisker, C.(2019) Mol Cell 74: 421
- PubMed: 30926243
- DOI: https://doi.org/10.1016/j.molcel.2019.02.029
- Primary Citation of Related Structures:
6H4H, 6H4I, 6H4J, 6H4K - PubMed Abstract:
Deubiquitinases have emerged as promising drug targets for cancer therapy. The two DUBs USP25 and USP28 share high similarity but vary in their cellular functions. USP28 is known for its tumor-promoting role, whereas USP25 is a regulator of the innate immune system and, recently, a role in tumorigenesis was proposed. We solved the structures of the catalytic domains of both proteins and established substantial differences in their activities. While USP28 is a constitutively active dimer, USP25 presents an auto-inhibited tetramer. Our data indicate that the activation of USP25 is not achieved through substrate or ubiquitin binding. USP25 cancer-associated mutations lead to activation in vitro and in vivo, thereby providing a functional link between auto-inhibition and the cancer-promoting role of the enzyme. Our work led to the identification of significant differences between USP25 and USP28 and provided the molecular basis for the development of new and highly specific anti-cancer drugs.
Organizational Affiliation:
Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, 97080 Würzburg, Germany.