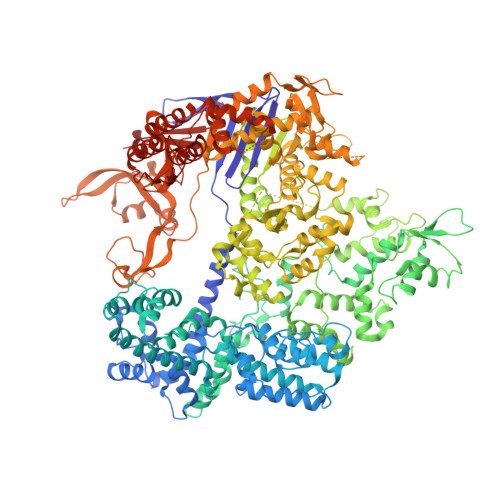
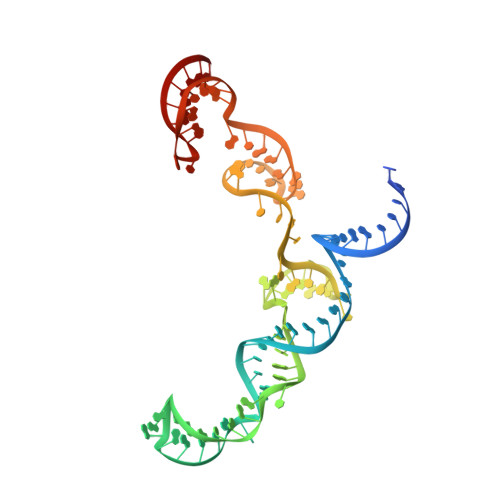
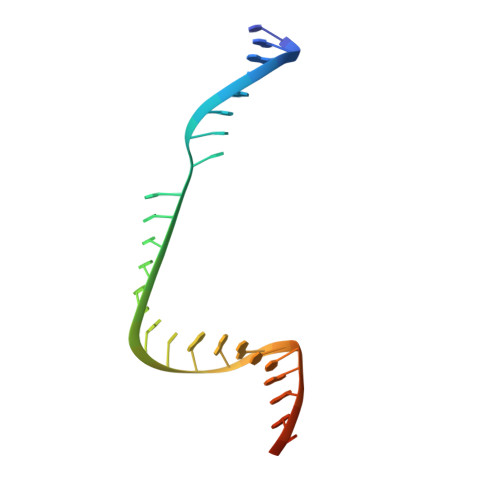
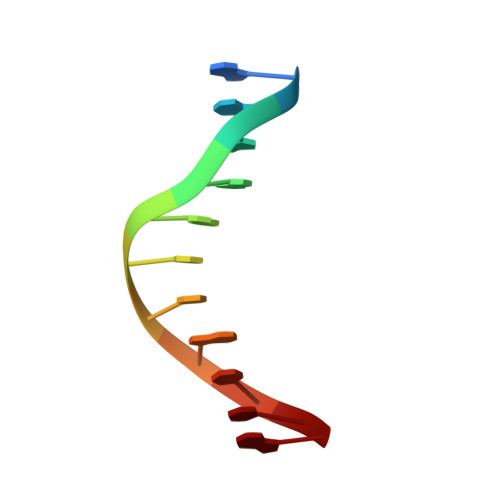
Structural insights into a high fidelity variant of SpCas9.
Guo, M., Ren, K., Zhu, Y., Tang, Z., Wang, Y., Zhang, B., Huang, Z.(2019) Cell Res 29: 183-192
- PubMed: 30664728
- DOI: https://doi.org/10.1038/s41422-018-0131-6
- Primary Citation of Related Structures:
6AEB, 6AEG - PubMed Abstract:
The RNA-guided endonucleases of the CRISPR-Cas9 system, including the most widely used Cas9 from Streptococcus pyogenes (SpCas9), are becoming a robust genome editing tool in model organisms and hold immense promise for therapeutic applications. Many strategies have been employed to overcome the limitations caused by SpCas9's off-target effects and its stringent requirement for the protospacer adjacent motif (PAM) sequence. However, the structural mechanisms underlying these strategies remain undefined. Here, we present crystal structure of a SpCas9 variant, xCas9 3.7 that has broad PAM compatibility and high DNA targeting specificity, in complex with a single-guide RNA and its double-stranded DNA targets. Structural comparison revealed that salt bridge-stabilized R1335 is critical for the stringent selection of PAM sequence by SpCas9. Unrestricted rotamerization of this residue by the E1219V mutation in xCas9 3.7 lessens the stringency for PAM recognition and allows SpCas9 to recognize multiple PAM sequences as further supported by biochemical data. Compared to those in wild-type (WT) SpCas9, REC2 and REC3 domains in xCas9 3.7 undergo striking conformational changes, leading to reduced contact with DNA substrate. SpCas9 mutants engineered to display less interaction with DNA and have conformationally more flexible REC2 and REC3 domains display enhanced specificity for DNA substrates in both biochemical and cellular assays. Taken together, our findings reveal the structural mechanisms underlying the broadened PAM compatibility and high DNA fidelity of xCas9 3.7, which can assist rational engineering of more efficient SpCas9 variants and probably other Cas9 orthologs.
Organizational Affiliation:
HIT Center for Life Sciences, School of Life Science and Technology, Harbin Institute of Technology, Harbin, Heilongjiang, 150080, China.