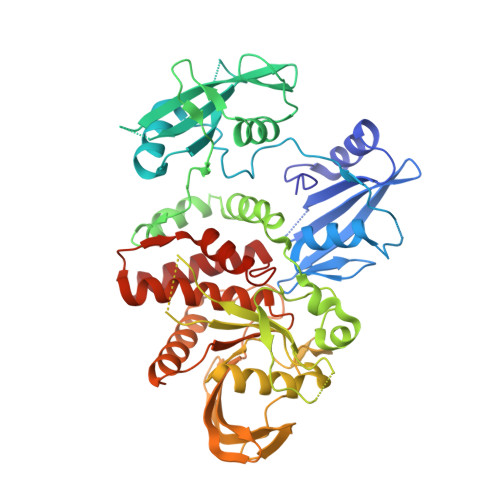
Structural and Functional Consequences of Three Cancer-Associated Mutations of the Oncogenic Phosphatase SHP2.
LaRochelle, J.R., Fodor, M., Xu, X., Durzynska, I., Fan, L., Stams, T., Chan, H.M., LaMarche, M.J., Chopra, R., Wang, P., Fortin, P.D., Acker, M.G., Blacklow, S.C.(2016) Biochemistry 55: 2269-2277
- PubMed: 27030275
- DOI: https://doi.org/10.1021/acs.biochem.5b01287
- Primary Citation of Related Structures:
5I6V, 5IBM, 5IBS - PubMed Abstract:
The proto-oncogene PTPN11 encodes a cytoplasmic protein tyrosine phosphatase, SHP2, which is required for normal development and sustained activation of the Ras-MAPK signaling pathway. Germline mutations in SHP2 cause developmental disorders, and somatic mutations have been identified in childhood and adult cancers and drive leukemia in mice. Despite our knowledge of the PTPN11 variations associated with pathology, the structural and functional consequences of many disease-associated mutants remain poorly understood. Here, we combine X-ray crystallography, small-angle X-ray scattering, and biochemistry to elucidate structural and mechanistic features of three cancer-associated SHP2 variants harboring single point mutations within the N-SH2:PTP interdomain autoinhibitory interface. Our findings directly compare the impact of each mutation on autoinhibition of the phosphatase and advance the development of structure-guided and mutation-specific SHP2 therapies.
Organizational Affiliation:
Department of Biological Chemistry & Molecular Pharmacology, Harvard Medical School , Boston, Massachusetts 02115, United States.