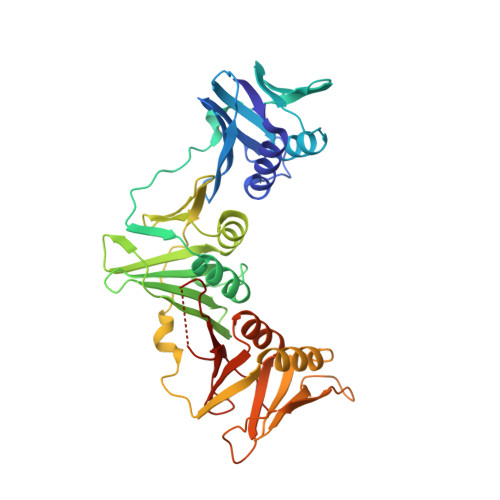
Targeting the beta-clamp in Helicobacter pylori with FDA-approved drugs reveals micromolar inhibition by diflunisal.
Pandey, P., Verma, V., Gautam, G., Kumari, N., Dhar, S.K., Gourinath, S.(2017) FEBS Lett 591: 2311-2322
- PubMed: 28656718
- DOI: https://doi.org/10.1002/1873-3468.12734
- Primary Citation of Related Structures:
5G48 - PubMed Abstract:
The β-clamp is the processivity-promoting factor for most of the enzymes in prokaryotic DNA replication; hence, it is a crucial drug target. In the present study, we investigated the β-clamp from Helicobacter pylori, aiming to seek potential drug molecules against this gastric-cancer-causing bacterium. An in silico screening of Food and Drug Administration (FDA) approved drugs against the H. pylori β-clamp, followed by its in vitro inhibition using a surface competition approach, yielded the drug diflunisal as a positive initial hit. Diflunisal inhibits the growth of H. pylori in the micromolar range. We determined the structure of diflunisal in complex with the β-clamp to show that the drug binds at subsite I, which is a protein-protein interaction site. Successful identification of FDA-approved molecules against H. pylori may lead to better and faster drug development.
Organizational Affiliation:
School of Life Sciences, Jawaharlal Nehru University, New Delhi, India.