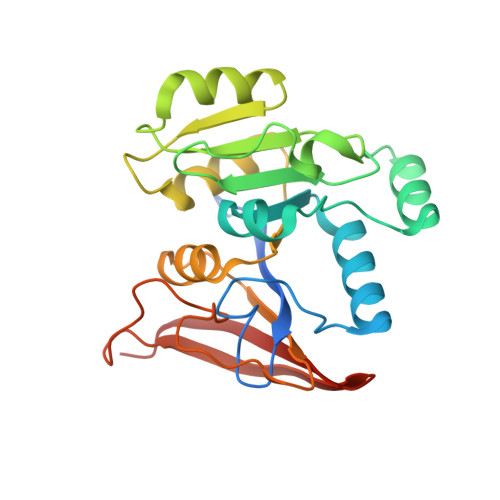
Evolution of archaeal Rib7 and eubacterial RibG reductases in riboflavin biosynthesis: Substrate specificity and cofactor preference.
Chen, S.C., Yen, T.M., Chang, T.H., Liaw, S.H.(2018) Biochem Biophys Res Commun 503: 195-201
- PubMed: 29864427
- DOI: https://doi.org/10.1016/j.bbrc.2018.06.002
- Primary Citation of Related Structures:
5XUX, 5XV0, 5XV2, 5XV5 - PubMed Abstract:
Archaeal/fungal Rib7 and eubacterial RibG possess a reductase domain for ribosyl reduction in the second and third steps, respectively, of riboflavin biosynthesis. These enzymes are specific for an amino and a carbonyl group of the pyrimidine ring, respectively. Here, several crystal structures of Methanosarcina mazei Rib7 are reported at 2.27-1.95 Å resolution, which are the first archaeal dimeric Rib7 structures. Mutational analysis displayed that no detectable activity was observed for the Bacillus subtilis RibG K151A, K151D, and K151E mutants, and the M. mazei Rib7 D33N, D33K, and E156Q variants, while 0.1-0.6% of the activity was detected for the M. mazei Rib7 N9A, S29A, D33A, and D57N variants. Our results suggest that Lys151 in B. subtilis RibG, while Asp33 together with Arg36 in M. mazei Rib7, ensure the specific substrate recognition. Unexpectedly, an endogenous NADPH cofactor is observed in M. mazei Rib7, in which the 2'-phosphate group interacts with Ser88, and Arg91. Replacement of Ser88 with glutamate eliminates the endogenous NADPH binding and switches preference to NADH. The lower melting temperature of ∼10 °C for the S88E and R91A mutants suggests that nature had evolved a tightly bound NADPH to greatly enhance the structural stability of archaeal Rib7.
Organizational Affiliation:
Structural Biology Program, National Yang-Ming University, No. 155, Sec. 2, Linong Street, Beitou District, Taipei, 11221, Taiwan.