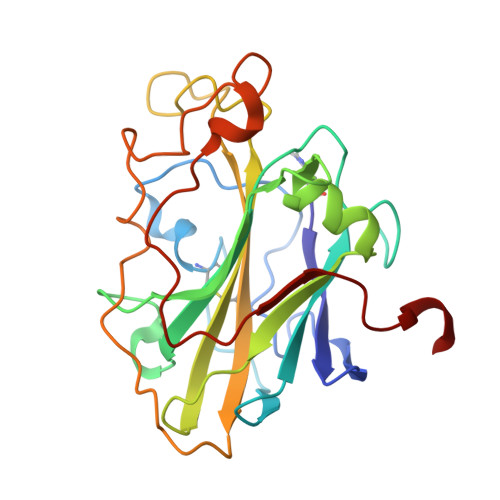
Structural and electronic determinants of lytic polysaccharide monooxygenase reactivity on polysaccharide substrates.
Simmons, T.J., Frandsen, K.E.H., Ciano, L., Tryfona, T., Lenfant, N., Poulsen, J.C., Wilson, L.F.L., Tandrup, T., Tovborg, M., Schnorr, K., Johansen, K.S., Henrissat, B., Walton, P.H., Lo Leggio, L., Dupree, P.(2017) Nat Commun 8: 1064-1064
- PubMed: 29057953
- DOI: https://doi.org/10.1038/s41467-017-01247-3
- Primary Citation of Related Structures:
5NKW, 5NLN, 5NLO, 5NLP, 5NLQ, 5NLR, 5NLS, 5NLT - PubMed Abstract:
Lytic polysaccharide monooxygenases (LPMOs) are industrially important copper-dependent enzymes that oxidatively cleave polysaccharides. Here we present a functional and structural characterization of two closely related AA9-family LPMOs from Lentinus similis (LsAA9A) and Collariella virescens (CvAA9A). LsAA9A and CvAA9A cleave a range of polysaccharides, including cellulose, xyloglucan, mixed-linkage glucan and glucomannan. LsAA9A additionally cleaves isolated xylan substrates. The structures of CvAA9A and of LsAA9A bound to cellulosic and non-cellulosic oligosaccharides provide insight into the molecular determinants of their specificity. Spectroscopic measurements reveal differences in copper co-ordination upon the binding of xylan and glucans. LsAA9A activity is less sensitive to the reducing agent potential when cleaving xylan, suggesting that distinct catalytic mechanisms exist for xylan and glucan cleavage. Overall, these data show that AA9 LPMOs can display different apparent substrate specificities dependent upon both productive protein-carbohydrate interactions across a binding surface and also electronic considerations at the copper active site.
Organizational Affiliation:
Department of Biochemistry, University of Cambridge, Cambridge, CB2 1QW, UK.