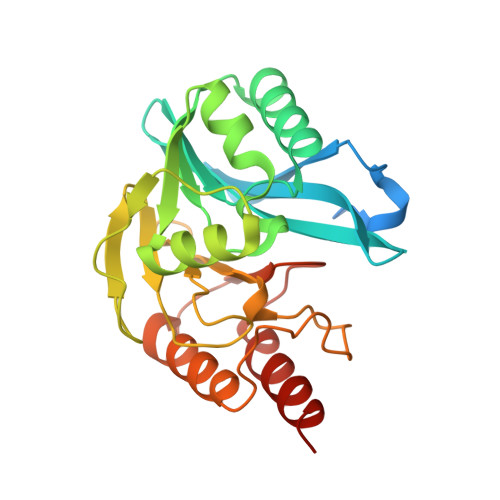
Metallo-beta-lactamase inhibitors by bioisosteric replacement: Preparation, activity and binding.
Skagseth, S., Akhter, S., Paulsen, M.H., Muhammad, Z., Lauksund, S., Leiros, H.S., Bayer, A.(2017) Eur J Med Chem 135: 159-173
- PubMed: 28445786
- DOI: https://doi.org/10.1016/j.ejmech.2017.04.035
- Primary Citation of Related Structures:
5MM9, 5NHZ, 5NI0 - PubMed Abstract:
Bacterial resistance is compromising the use of β-lactam antibiotics including carbapenems. The main resistance mechanism against β-lactams is hydrolysis of the β-lactam ring mediated by serine- or metallo-β-lactamases (MBLs). Although several inhibitors of MBLs have been reported, none has been developed into a clinically useful inhibitor. Mercaptocarboxylic acids are among the most prominent scaffolds reported as MBL inhibitors. In this study, the carboxylate group of mercaptocarboxylic acids was replaced with bioisosteric groups like phosphonate esters, phosphonic acids and NH-tetrazoles. The influence of the replacement on the bioactivity and inhibitor binding was evaluated. A series of bioisosteres of previously reported inhibitors was synthesized and evaluated against the MBLs VIM-2, NDM-1 and GIM-1. The most active inhibitors combined a mercapto group and a phosphonate ester or acid, with two/three carbon chains connecting a phenyl group. Surprisingly, also compounds containing thioacetate groups instead of thiols showed low IC 50 values. High-resolution crystal structures of three inhibitors in complex with VIM-2 revealed hydrophobic interactions for the diethyl groups in the phosphonate ester (inhibitor 2b), the mercapto bridging the two active site zinc ions, and tight stacking of the benzene ring to the inhibitor between Phe62, Tyr67, Arg228 and His263. The inhibitors show reduced enzyme activity in Escherichia coli cells harboring MBL. The obtained results will be useful for further structural guided design of MBL inhibitors.
Organizational Affiliation:
The Norwegian Structural Biology Centre (NorStruct), Department of Chemistry, Faculty of Science and Technology, UiT The Arctic University of Norway, N-9037 Tromsø, Norway.