X-ray structure of the H235Q mutant of GLIC in complex with propofol
Fourati, Z., Delarue, M.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Proton-gated ion channel | 327 | Gloeobacter violaceus PCC 7421 | Mutation(s): 1 Gene Names: glvI, glr4197 Membrane Entity: Yes | 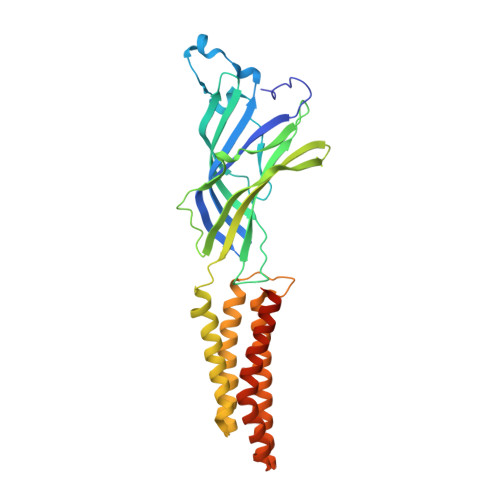 | |
UniProt | |||||
Find proteins for Q7NDN8 (Gloeobacter violaceus (strain ATCC 29082 / PCC 7421)) Explore Q7NDN8 Go to UniProtKB: Q7NDN8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7NDN8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLC Query on PLC | AA [auth C] BA [auth C] CA [auth C] F [auth A] G [auth A] | DIUNDECYL PHOSPHATIDYL CHOLINE C32 H65 N O8 P IJFVSSZAOYLHEE-SSEXGKCCSA-O |  | ||
| LMT Query on LMT | GA [auth C] N [auth A] O [auth A] PA [auth D] Y [auth B] | DODECYL-BETA-D-MALTOSIDE C24 H46 O11 NLEBIOOXCVAHBD-QKMCSOCLSA-N |  | ||
| PFL Query on PFL | AB [auth E], HA [auth C], Q [auth A], RA [auth D] | 2,6-BIS(1-METHYLETHYL)PHENOL C12 H18 O OLBCVFGFOZPWHH-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | FA [auth C] IA [auth D] M [auth A] OA [auth D] P [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CL Query on CL | DA [auth C] I [auth A] J [auth A] K [auth A] MA [auth D] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Query on NA | EA [auth C] L [auth A] NA [auth D] R [auth B] W [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 182.842 | α = 90 |
| b = 132.989 | β = 102.83 |
| c = 161.162 | γ = 90 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| REFMAC | phasing |