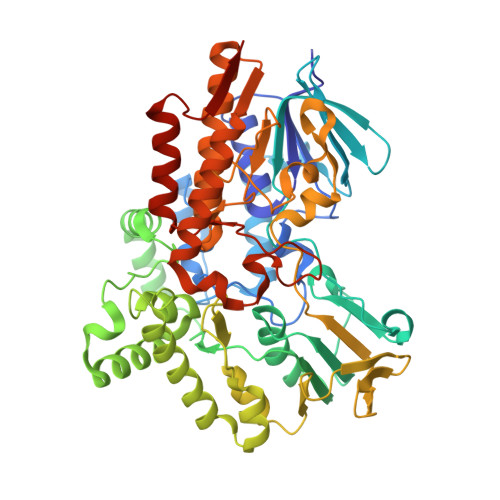
Characterization and Crystal Structure of a Robust Cyclohexanone Monooxygenase.
Romero, E., Castellanos, J.R., Mattevi, A., Fraaije, M.W.(2016) Angew Chem Int Ed Engl 55: 15852-15855
- PubMed: 27873437
- DOI: https://doi.org/10.1002/anie.201608951
- Primary Citation of Related Structures:
5M0Z, 5M10 - PubMed Abstract:
Cyclohexanone monooxygenase (CHMO) is a promising biocatalyst for industrial reactions owing to its broad substrate spectrum and excellent regio-, chemo-, and enantioselectivity. However, the low stability of many Baeyer-Villiger monooxygenases is an obstacle for their exploitation in industry. Characterization and crystal structure determination of a robust CHMO from Thermocrispum municipale is reported. The enzyme efficiently converts a variety of aliphatic, aromatic, and cyclic ketones, as well as prochiral sulfides. A compact substrate-binding cavity explains its preference for small rather than bulky substrates. Small-scale conversions with either purified enzyme or whole cells demonstrated the remarkable properties of this newly discovered CHMO. The exceptional solvent tolerance and thermostability make the enzyme very attractive for biotechnology.
Organizational Affiliation:
Department of Biotechnology, University of Groningen, Nijenborgh 4, 9747AG, Groningen, The Netherlands.