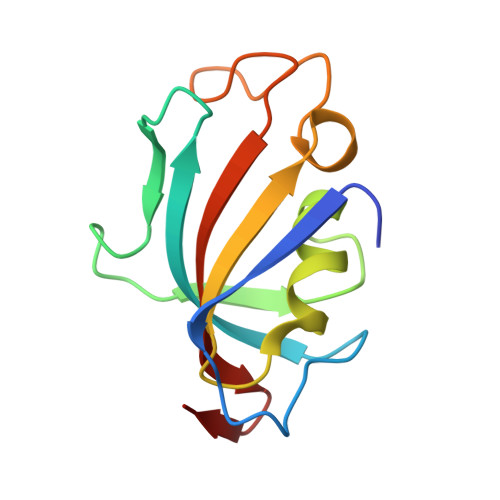
Structures of Pathogenic Fungal FKBP12s Reveal Possible Self-Catalysis Function.
Tonthat, N.K., Juvvadi, P.R., Zhang, H., Lee, S.C., Venters, R., Spicer, L., Steinbach, W.J., Heitman, J., Schumacher, M.A.(2016) mBio 7: e00492-e00416
- PubMed: 27118592
- DOI: https://doi.org/10.1128/mBio.00492-16
- Primary Citation of Related Structures:
5HT1, 5HTG, 5HUA, 5HW6, 5HW7, 5HW8, 5HWB, 5HWC, 5I98, 5J6E - PubMed Abstract:
Invasive fungal infections remain difficult to treat and require novel targeting strategies. The 12-kDa FK506-binding protein (FKBP12) is a ubiquitously expressed peptidyl-prolyl isomerase with considerable homology between fungal pathogens and is thus a prime candidate for future targeting efforts to generate a panfungal strategy. Despite decades of research on FKBPs, their substrates and mechanisms of action remain unclear. Here we describe structural, biochemical, and in vivo analyses of FKBP12s from the pathogenic fungi Candida albicans, Candida glabrata, and Aspergillus fumigatus Strikingly, multiple apo A. fumigatus and C. albicans FKBP12 crystal structures revealed a symmetric, intermolecular interaction involving the deep insertion of an active-site loop proline into the active-site pocket of an adjacent subunit. Such interactions have not been observed in previous FKBP structures. This finding indicates the possibility that this is a self-substrate interaction unique to the A. fumigatus and C. albicans fungal proteins that contain this central proline. Structures obtained with the proline in the cis and trans states provide more data in support of self-catalysis. Moreover, cysteine cross-linking experiments captured the interacting dimer, supporting the idea that it forms in solution. Finally, genetic studies exploring the impact of mutations altering the central proline and an adjacent residue provide evidence that any dimeric state formed in vivo, where FKBP12 concentrations are low, is transient. Taken together, these findings suggest a unique mechanism of self-substrate regulation by fungal FKBP12s, lending further novel understanding of this protein for future drug-targeting efforts.
Organizational Affiliation:
Department of Biochemistry, Duke University School of Medicine, Durham, North Carolina, USA.