Crystal Structure of Jmjc Domain of Human Histone Demethylase Uty in Complex with Succinate
Nowak, R., Krojer, T., Johansson, C., Gileadi, C., Kupinska, K., von Delft, F., Arrowsmith, C.H., Bountra, C., Edwards, A., Oppermann, U.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HISTONE DEMETHYLASE UTY | 478 | Homo sapiens | Mutation(s): 0 | 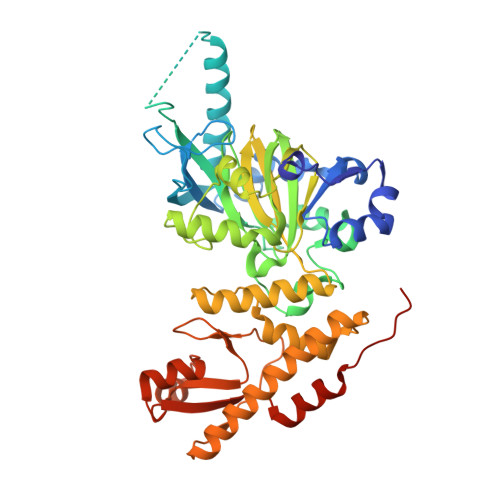 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for O14607 (Homo sapiens) Explore O14607 Go to UniProtKB: O14607 | |||||
PHAROS: O14607 GTEx: ENSG00000183878 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O14607 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SIN Query on SIN | D [auth A], N [auth B] | SUCCINIC ACID C4 H6 O4 KDYFGRWQOYBRFD-UHFFFAOYSA-N |  | ||
| ZN Query on ZN | L [auth A], X [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | E [auth A] F [auth A] G [auth A] H [auth A] I [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| MN Query on MN | C [auth A], M [auth B] | MANGANESE (II) ION Mn WAEMQWOKJMHJLA-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 91.69 | α = 90 |
| b = 110.77 | β = 90 |
| c = 119.93 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| SCALA | data scaling |
| DIMPLE | phasing |