Design and evaluation of novel glutaminase inhibitors.
McDermott, L.A., Iyer, P., Vernetti, L., Rimer, S., Sun, J., Boby, M., Yang, T., Fioravanti, M., O'Neill, J., Wang, L., Drakes, D., Katt, W., Huang, Q., Cerione, R.(2016) Bioorg Med Chem 24: 1819-1839
- PubMed: 26988803
- DOI: https://doi.org/10.1016/j.bmc.2016.03.009
- Primary Citation of Related Structures:
5FI2, 5FI6, 5FI7, 5I94 - PubMed Abstract:
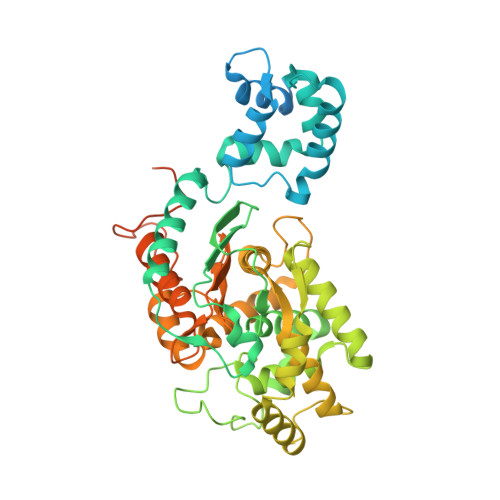
A novel set of GAC (kidney glutaminase isoform C) inhibitors able to inhibit the enzymatic activity of GAC and the growth of the triple negative MDA-MB-231 breast cancer cells with low nanomolar potency is described. Compounds in this series have a reduced number of rotatable bonds, improved ClogPs, microsomal stability and ligand efficiency when compared to the leading GAC inhibitors BPTES and CB-839. Property improvements were achieved by the replacement of the flexible n-diethylthio or the n-butyl moiety present in the leading inhibitors by heteroatom substituted heterocycloalkanes.
Organizational Affiliation:
University of Pittsburgh, Department of Pharmaceutical Sciences, Pittsburgh, PA 15261, USA; University of Pittsburgh, Drug Discovery Institute, Pittsburgh, PA 15261, USA. Electronic address: lam179@pitt.edu.